fnr:On One Page
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;} h2 .editsection { display:none;}</css>
| Quickview | Gene | Gene Product(s) | Expression | Evolution | On One Page |
| Quickview | | Gene | | Product(s) | | Expression| | Evolution | | References | |
Quickview
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
| Standard Name |
fnr |
|---|---|
| Gene Synonym(s) | |
| Product Desc. |
FNR transcriptional dual regulator[2][3] Global transcription factor for anaerobic growth; [4FE-4S] cluster; homodimeric[4] |
| Product Synonyms(s) |
DNA-binding transcriptional dual regulator, global regulator of anaerobic growth[1], B1334[2][1], Fnr[2][1], OssA[2][1], NirR[2][1], NirA[2][1], transcriptional regulation of aerobic, anaerobic respiration, osmotic balance[2][1], OxrA[2][1] , ECK1330, JW1328, nirA, nirR, ossA, b1334 |
| Function from GO |
<GO_nr /> |
| Knock-Out Phenotype | |
| Regulation/Expression | |
| Regulation/Activity | |
| Quick Links | |
| edit table |
</protect> See Help:Quickview for help with entering information in the Quickview table. <protect></protect>
Notes
FNR represses genes involved in aerobic respiration and activates genes required for anaerobic respiration.[4]
Gene
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
Nomenclature
See Help:Gene_nomenclature for help with entering information in the Gene Nomenclature table.
| Standard name |
fnr |
|---|---|
| Mnemonic |
fumurate and nitrate reductase Fumarate and nitrate reduction |
| Synonyms | |
| edit table |
</protect>
Notes
The fnr gene was identified in 1976 by Lambden and Guest [5] who isolated mutants of E. coli K-12 that were unable to use fumarate as an anaerobic electron acceptor.
Location(s) and DNA Sequence
<protect>
See Help:Gene_location for help entering information in the Gene Location and DNA sequence table.
| Strain | Map location | Genome coordinates | Genome browsers | Sequence links |
|---|---|---|---|---|
|
MG1655 |
30.11 minutes |
MG1655: 1397550..1396798 |
||
|
NC_012967: 1396696..1395944 |
||||
|
NC_012759: 1287647..1288399 |
||||
|
W3110 |
|
W3110: 1401240..1400488 |
||
|
DH10B: 1486946..1486194 |
||||
| edit table |
</protect>
Notes
Sequence Features
See Help:Gene_sequence_features for help in entering sequence features in EcoliWiki.
| Feature Type | Strain | Genomic Location | Evidence | Reference | Notes |
|---|---|---|---|---|---|
|
Coding Start (SO:0000323) |
1396798 |
Edman degradation |
PMID:2181237 |
| |
| edit table |
<protect></protect>
Notes
Alleles and Phenotypes
See Help:Gene_alleles for how to enter or edit alleles and phenotypes in EcoliWiki.
| Allele | Nt change(s) | AA change(s) | Phenotype: Type | Phenotype: Description | Reference | Availability | Comments |
|---|---|---|---|---|---|---|---|
|
fnr(del) (Keio:JW1328) |
deletion |
deletion |
PMID:16738554 |
||||
|
fnrD154A,G,V |
D154A,G,V |
Loss of regulation by O(2) |
seeded from UniProt:P0A9E5 | ||||
|
fnr::Tn5KAN-I-SceI (FB20296) |
Insertion at nt 606 in Plus orientation |
PMID:15262929 |
does not contain pKD46 | ||||
|
fnrC16A |
C16A |
No effect |
PMID:2226775 |
published as fnr-382 seeded from UniProt:P0A9E5 | |||
|
fnrC20S |
C20S |
Loss of activity |
PMID:2226775 |
published as fnr-197 seeded from UniProt:P0A9E5 | |||
|
fnrD22G |
D22G |
Loss of regulation by O(2) |
seeded from UniProt:P0A9E5 | ||||
|
fnrS73F |
S73F |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrE64Q |
E64Q |
No effect |
seeded from UniProt:P0A9E5 | ||||
|
fnrR72H |
R72H |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrI81T |
I81T |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrG85A |
G85A |
Decrease in activity; no effect on DNA-binding. Trace activity; when associated with P-187 |
seeded from UniProt:P0A9E5 | ||||
|
fnrD86A |
D86A |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrE87K |
E87K |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrQ88E |
Q88E |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrH93R |
H93R |
Loss of regulation by O(2) |
seeded from UniProt:P0A9E5 | ||||
|
fnrG96D |
G96D |
Loss of activity |
seeded from UniProt:P0A9E5 | ||||
|
fnrT118A,P |
T118A,P |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrM120I,R,T,V |
M120I,R,T,V |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrC122A,S |
C122A,S |
Loss of activity |
seeded from UniProt:P0A9E5 | ||||
|
fnrR140A |
R140A |
Decrease in activity |
seeded from UniProt:P0A9E5 | ||||
|
fnrM143A |
M143A |
Decrease in activity |
seeded from UniProt:P0A9E5 | ||||
|
fnrM144A |
M144A |
Decrease in activity due to faulty dimerization |
seeded from UniProt:P0A9E5 | ||||
|
fnrR145A |
R145A |
Decrease in activity |
seeded from UniProt:P0A9E5 | ||||
|
fnrL146A |
L146A |
Decrease in activity |
seeded from UniProt:P0A9E5 | ||||
|
fnrM147A |
M147A |
Decrease in activity due to faulty dimerization |
seeded from UniProt:P0A9E5 | ||||
|
fnrE150K |
E150K |
Loss of regulation by O(2) |
seeded from UniProt:P0A9E5 | ||||
|
fnrT82P |
T82P |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrI151A |
I151A |
Decrease in activity due to faulty dimerization |
seeded from UniProt:P0A9E5 | ||||
|
fnrC29G |
C29G |
Loss of activity |
seeded from UniProt:P0A9E5 | ||||
|
fnrD43G |
D43G |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrI158A |
I158A |
Decrease in activity due to faulty dimerization |
seeded from UniProt:P0A9E5 | ||||
|
fnrF181L |
F181L |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrF186S |
F186S |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrS187P |
S187P |
Decrease in activity; no effect on DNA-binding. Trace activity; when associated with A-85 |
seeded from UniProt:P0A9E5 | ||||
|
fnrF191L |
F191L |
Decrease in activity; no effect on DNA-binding |
seeded from UniProt:P0A9E5 | ||||
|
fnrC23G |
C23G |
Loss of activity |
PMID:2226775 |
published as fnr-383 seeded from UniProt:P0A9E5 | |||
|
fnrL28H |
L28H |
Loss of regulation by O(2) |
seeded from UniProt:P0A9E5 | ||||
|
fnr(del)::kan |
deletion |
Biolog:respiration |
unable to respire L-Asparagine |
PMID:16095938 |
|||
|
D154A |
D154A |
Constitutive dimerization |
PMID:16959764 |
||||
|
fnr-1 |
|||||||
|
fnr-25 |
|||||||
|
fnr-267(del) |
|||||||
|
fnr-771(del)::kan |
PMID:16738554 |
||||||
|
fnr - |
Sensitivity to |
Mutants were sensitive to minimun 5 mM of chlorate. |
PMID:7047497 |
| |||
| edit table |
<protect></protect>
Notes
Genetic Interactions
<protect>
| Interactor | Interaction | Allele | Score(s) | Reference(s) | Accessions | Notes |
|---|---|---|---|---|---|---|
| edit table |
</protect>
Notes
Genetic Resources
See Help:Gene_resources for help entering information into the Genetic Resources table.
| Resource | Resource Type | Source | Notes/Reference |
|---|---|---|---|
|
JW1328 |
Plasmid clone |
PMID:16769691 Status:Clone OK Primer 1:GCCATCCCGGAAAAGCGAATTAT Primer 2:CCGGCAACGTTACGCGTATGACC | |
|
Kohara Phage |
PMID:3038334 | ||
|
Kohara Phage |
PMID:3038334 | ||
|
Linked marker |
est. P1 cotransduction: 34% [7] | ||
|
Linked marker |
est. P1 cotransduction: 62% [7] | ||
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG10325 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10325 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120000319 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EchoBASE:EB0321 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0004475 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Product(s)
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Nomenclature
See Help:Product_nomenclature for help entering or editing information in this section of EcoliWiki.
| Standard name |
Fnr |
|---|---|
| Synonyms |
DNA-binding transcriptional dual regulator, global regulator of anaerobic growth[1], B1334[2][1], Fnr[2][1], OssA[2][1], NirR[2][1], NirA[2][1], transcriptional regulation of aerobic, anaerobic respiration, osmotic balance[2][1], OxrA[2][1] , ECK1330, JW1328, nirA, nirR, ossA, b1334 |
| Product description |
FNR transcriptional dual regulator[2][3] Global transcription factor for anaerobic growth; [4FE-4S] cluster; homodimeric[4] |
| EC number (for enzymes) |
|
| edit table |
<protect></protect>
Notes
FNR is a global regulator, belonging to the CRP superfamily of helix turn helix DNA binding proteins. FNR is active under anaerobic conditions and depending on the location of the binding site, either activates or represses genes. The oxygen sensing function of FNR is mediated by a [4Fe-4S] cluster. The [4Fe-4S] cluster is ligated to Cys20, 23, 29 and 122, is stable anaerobically and promotes FNR dimerization. In the presence of oxygen, the cluster is converted to a [2Fe-2S] cluster decreasing FNR dimerization, DNA binding and inactivating FNR.
Function
<protect>
Gene Ontology
See Help:Gene_ontology for help entering or editing GO terms and GO annotations in EcoliWiki.
| Qualifier | GO ID | GO term name | Reference | Evidence Code | with/from | Aspect | Notes | Status |
|---|---|---|---|---|---|---|---|---|
|
GO:0003677 |
DNA binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012318 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003677 |
DNA binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0238 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003700 |
transcription factor activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR001808 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003700 |
transcription factor activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR018335 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005506 |
iron ion binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0408 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005622 |
intracellular |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR001808 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005622 |
intracellular |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR018335 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005737 |
cytoplasm |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0963 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005737 |
cytoplasm |
GO_REF:0000023 |
IEA: Inferred from Electronic Annotation |
SP_SL:SL-0086 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006350 |
transcription |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0804 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006355 |
regulation of transcription, DNA-dependent |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR001808 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006355 |
regulation of transcription, DNA-dependent |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR018335 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0045449 |
regulation of transcription |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012318 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0045449 |
regulation of transcription |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0805 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0046872 |
metal ion binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0479 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0051536 |
iron-sulfur cluster binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0411 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0051539 |
4 iron, 4 sulfur cluster binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0004 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
| edit table |
Interactions See Help:Product_interactions for help entering or editing information about gene product interactions in this section of EcoliWiki.
| Partner Type | Partner | Notes | References | Evidence |
|---|---|---|---|---|
|
Protein |
phoB |
PMID:16606699 |
Experiment(s):EBI-1139571 | |
|
Protein |
nadE |
PMID:16606699 |
Experiment(s):EBI-1139571 | |
|
Protein |
prs |
PMID:16606699 |
Experiment(s):EBI-1139571 | |
|
Protein |
matC |
PMID:16606699 |
Experiment(s):EBI-1139571 | |
|
Protein |
self |
detectable dimerization requires ligation of the [4Fe-4S] cluster |
| |
| edit table |
</protect>
Notes
Localization
See Help:Product_localization for how to add or edit information in this section of EcoliWiki.
| Compartment | Description | Evidence | Reference/Source | Notes |
|---|---|---|---|---|
|
cytoplasm |
From EcoCyc[3] |
| ||
| edit table |
<protect></protect>
Notes
Structure and Physical Properties
<protect>
Physical Properties
See Help:Product_physical_properties for help entering or editing information about the physical properties of this gene product.
| Name | |
|---|---|
| Sequence |
MIPEKRIIRR IQSGGCAIHC QDCSISQLCI PFTLNEHELD QLDNIIERKK PIQKGQTLFK AGDELKSLYA IRSGTIKSYT ITEQGDEQIT GFHLAGDLVG FDAIGSGHHP SFAQALETSM VCEIPFETLD DLSGKMPNLR QQMMRLMSGE IKGDQDMILL LSKKNAEERL AAFIYNLSRR FAQRGFSPRE FRLTMTRGDI GNYLGLTVET ISRLLGRFQK SGMLAVKGKY ITIENNDALA QLAGHTRNVA |
| Length |
250 |
| Mol. Wt |
27.968 kDa |
| pI |
8.1 (calculated) |
| Extinction coefficient |
7,450 - 8,075 (calc based on 5 Y, 0 W, and 5 C residues) |
| edit table |
Domains/Motifs/Modification Sites
|
See Help:Product_domains_motifs for help entering or editing information in this section of EcoliWiki.
|
<motif_map/> |
|
Structure
| </protect>
Structure figures<protect> | ||||||
Notes
Gene Product Resources
See Help:Product_resources for help with entering or editing information in this section of EcoliWiki.
| Resource type | Source | Notes/Reference |
|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
ASAP:ABE-0004475 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EcoCyc:EG10325 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10325 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
RegulonDB:ECK120000319 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB0321 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Expression
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Overview
This is a placeholder for a summary statement on how expression of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Cellular Levels
| Molecule | Organism or Strain | Value | Units | Experimental Conditions | Assay used | Notes | Reference(s) |
|---|---|---|---|---|---|---|---|
|
Protein |
E. coli K-12 MC4100 |
4.53E+02 |
molecules/cell |
|
emPAI |
PMID:18304323 | |
|
Protein |
E. coli K-12 MG1655 |
6551 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
2582 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
2504 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
| edit table |
Notes
Transcription and Transcriptional Regulation
<protect>
|
See Help:Expression_transcription for help entering or editing information in this section of EcoliWiki.
|
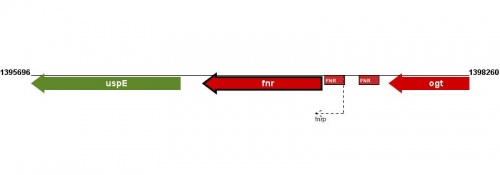 Figure courtesy of RegulonDB |
</protect>
Notes
This is a placeholder for a summary statement about how transcription of this gene is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Translation and Regulation of Translation
<protect><gbrowseImage>
name=NC_000913:1397530..1397570
source=MG1655
flip=1
type=Gene+DNA_+Protein
preset=Nterminus
</gbrowseImage>
This picture shows the sequence around the N-terminus.
</protect>
Notes
This is a placeholder for a summary statement about how translation of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Turnover and Regulation of Turnover
</protect>
Notes
This is a placeholder for a summary statement about turnover of this gene product. You can help EcoliWiki by becoming a user and writing/editing this statement.
Experimental
<protect>
Mutations Affecting Expression
See Help:Expression_mutations for help entering or editing information in this section of EcoliWiki.
| Allele Name | Mutation | Phenotype | Reference |
|---|---|---|---|
| edit table |
Expression Studies
See Help:Expression_studies for help entering or editing information in this section of EcoliWiki.
| Type | Reference | Notes |
|---|---|---|
|
microarray |
NCBI GEO profiles for fnr | |
|
microarray |
Summary of data for fnr from multiple microarray studies | |
| edit table |
Expression Resources
See Help:Expression_resources for help entering or editing information in this section of EcoliWiki.
| Resource Name | Resource Type | Description | Source | Notes |
|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions Related to fnr Expression
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG10325 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB0321 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10325 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120000319 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0004475 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Evolution
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Homologs in Other Organisms
See Help:Evolution_homologs for help entering or editing information in this section of EcoliWiki.
| Organism | Homologs (Statistics) | Comments |
|---|---|---|
|
Homo sapiens |
|
From Inparanoid:20070104 |
|
Xenopus tropicalis |
|
From Inparanoid:20070104 |
|
Shigella flexneri |
FNR |
From SHIGELLACYC |
|
E. coli O157 |
FNR |
From ECOO157CYC |
| edit table |
Do-It-Yourself Web Tools
<protect></protect>
Notes
Families
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG10325 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10325 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120000319 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB0321 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0004475 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
References
See Help:References for how to manage references in EcoliWiki.
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 1.17 Riley, M. et al. (2006) Nucleic Acids Res 34:1-6 (corrected supplemental data from B. Wanner)
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 2.11 2.12 2.13 2.14 2.15 2.16 2.17 2.18 EcoCyc (release 10.6; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 3.0 3.1 3.2 EcoCyc (release 11.1; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 4.0 4.1 4.2 EcoGene: Rudd, KE (2000) EcoGene: a genome sequence database for Escherichia coli K-12. Nucleic Acids Res 28:60-4.
- ↑ Lambden, PR & Guest, JR (1976) Mutants of Escherichia coli K12 unable to use fumarate as an anaerobic electron acceptor. J. Gen. Microbiol. 97 145-60 PubMed
- ↑ 6.0 6.1 6.2 CGSC: The Coli Genetics Stock Center
- ↑ 7.0 7.1 The Tn10 insertion sites determined by Nichols et al. 1998 (PMID:9829956) were reannotated by alignment with E. coli K-12 genome sequence (GenBank accession NC_000913). P1 contransduction frequencies were calculated using the formula of Wu (PMID:5338813).
Categories


