Help:Gene Ontology
The Gene Ontology (GO) is a collaborative effort to "standardize the representation of gene and gene product attributes across species and databases." [1]
Overview
The GO can be divided into three main areas:
- a controlled vocabulary of terms for the products of genes
- a set of relationships between those terms such as (such as "is a", or "part of")
- annotations about specific gene products from various species
Ecoliwiki would like to capture as much information as possible in a way that is the most useful to other scientists. GO helps accomplish that task. On each Gene Product page there is a table for GO annotations. The community encourages the addition of any information you could provide. Even if you are not familiar with GO you can contribute, someone else can fill in the missing details later.
GO table
The GO tables on each of the Gene Product pages contain all the GO annotations for that particular product. There are nine columns, most of the time only 6 or 7 of which are used. A quick overview of each of the columns is as such:
- Qualifier
- This is a term that is used to modify the interpretation of an annotation. For example an annotation that was found to be incorrect can have a "NOT" added as the qualifier.
- Some qualifiers used in EcoliWiki are: NOT, Under_review, and Deprecated
- GO ID
- The unique number associated with each term. (Prefixed with "GO: ". )
- GONUTS wiki can help you find a suitable GO term.
- GO term name
- The name of the term. Here are some example terms:
- Reference(s)
- The reference for this annotation. Typically a Pubmed article ID (prefixed with a "PMID: ".)
- Evidence Code
- Probably the most complicated of the fields, the evidence code specifies how the information for this annotation was gained.
- Some examples are "IDA: Inferred from Direct Assay" and "ISS: Inferred from Sequence or Structural Similarity".
- A more complete guide with explanations and some examples of the evidence codes can be found here .
- with/from
- Used when a comparative annotation is used.http://www.geneontology.org/GO.evidence.shtml
- For example, when the evidence is "ISS: Inferred from Sequence or Structural Similiarity" is used, you must specify which sequence/structure it is similar to. (Prefixed with a database prefix such as "PDB:" or "UniProt:")
- Other evidence codes may require a protein partner, etc. See the Guide mentioned above.
- Aspect
- A single letter to describe the specific ontology this term is in.
- This is not required by the user, and will be filled in automatically when you enter a GO ID.
- Notes
- A free-text place for user notes, big or small.
- Status
- If this annotation has the required three things: GO ID, reference, and evidence it is complete.
To see a sample page, have a look at the product page for dnaJ.
Editing and Entering Information
Editing and entering information into the tables is relatively simple. Please note that the titles of the boxes in the pictures, and the text in the boxes, may be different from the table you are editing.
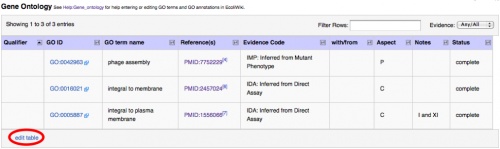
To add new data or to modify existing data, click the edit table link on the bottom, left-hand side of the Gene Ontology table. An editable version of the table will appear. An example is shown in the next image.
There are a couple options that let you add new data:
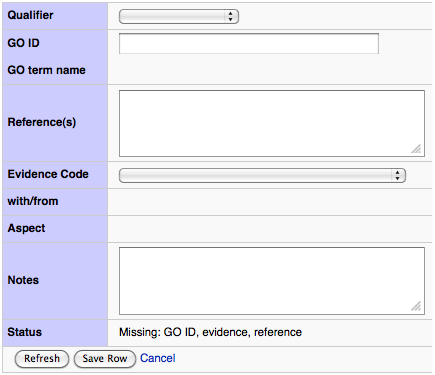
- The best (and easiest!) is the Add row button near the bottom of the table. This will take you to the form shown in the next image. It is strongly recommended that you use this option.
- The Add multiple button allows several rows to be added simultaneously, but does not automatically add information like the form (eg, the "GO term name") nor does it verify you made a complete entry (see next caption for more details). If you know all the information needed and wish to use this function, the syntax is provided on the page you enter your data on. If you are using the Add multiple function and you need to leave a column blank (eg, neither your GO term nor your evidence require a with/from entry), do not enter any characters between the pipes.
Several fields that are editable from this page, and several are filled in based on the rest of the information in the table. A quality annotation needs at least a GO ID, a Reference, and an Evidence Code. If you only have enough information to add a partial annotation, you are encouraged to do so; it will not be marked "complete" but it is expected the community will update your annotation as more information is made available.
- The Qualifier field is a drop-down menu that is only applicable in a few cases, and is generally left blank.
- The GO ID field is required. Enter the identification number corresponding to the desired GO term in the format GO:XXXXXXX. You can just copy and paste this from the term's page.
- The GO term name will be automatically filled in by the computer, and helps you verify that you used the right GO ID in the previous field.
- The Reference(s) field is also required, and will add a link to Ecoliwiki's page on that paper if you enter the PubMed(PubMed on EcoliWiki) identification number, PMID, in the format PMID:XXXXXXXX. You may also use a PMCID (PubMed Central Identification) if necessary, but try using the converter at http://www.ncbi.nlm.nih.gov/sites/pmctopmid first. On rare occasions other non-journal articles may be cited (theses, books, etc); please try to give as much identification as you can in order to allow others to easily find the information. DOIs may also be used but PMIDs are the strongly preferred identification- please use if one is available. Typically only one reference is needed per annotation. If you have multiple options choose the oldest source that supports the most detailed GO term available. If you have references other than a PMID, see the References Help page for help formatting your sources.
- The Evidence Code drop-down menu has the available evidence codes, and this field is mandatory for a "complete" annotation. More information, and examples, can be found at http://www.geneontology.org/GO.evidence.shtml. Not all evidence codes shown in the guide may be available to all users, but all except the rarest evidence codes are available to all users.
- The with/from field will open up if an evidence code that can or must use the field is selected. Depending on the requirements of the evidence code, a Uniprot accession number, GO ID, or several other types of IDs can be entered. Select the appropriate field from the drop-down menu and fill in the field. Note: the following example IDs are completely random, but links are provided so you can become familiar with the assigning website.
- EcoliWiki: needs the four-letter name assigned by Ecoliwiki; use the exact capitalization on that gene's page.
- UniProtKB: use the UniProt accession number; this will usually be alphanumeric and resemble P03040.
- InterPro: this can be found at the EBI website and should be in the form IPR004091.
- EcoCyc: EcoCyc accession numbers are also alphanumeric and are in the form EG10560.
- PMID: a document's numeric PubMed identification in the form 6981459 can be found by searching the PubMed site.
- Another row will appear once one is filled in in case multiple rows are needed (eg, an IGI that needs three proteins entered in the with/from field will use three rows of information in this area).
- The Aspect field will be filled out automatically when you enter a GO ID. This field simply indicates if the GO ID is a Process, Function, or Component term.
- Although the Notes field is optional, clarifications or comments applicable to the GO and the gene/protein may be entered here. The paper itself has its own EcoliWiki page, and comments pertaining to the whole paper or more than the specific gene/protein may be entered there.
- The Status field is another uneditable field, and will tell you if the annotation is complete, or if it isn't complete it will list which of the mandatory fields you still need to complete. This field will not tell you if your entries are in the recommended format.
After entering all the information you can, or making a complete entry, the Refresh button will reload the Go term name, Status, and other automatic columns. This is especially useful when editing an entry, as it ensures the GO term name matches the GO ID (if you changed it). The Status may not also not reflect recent changes unless you refresh before saving. The Refresh button is not required, and annotations can be made without using it, but it is recommended as the most recent Status verifies you have included all the components of a complete annotation. Note: Neither the Refresh button, nor a status of "complete", are indications that an annotation is correct, merely that all components are present and in the most recent form.
There may be information already in the table that you wish to change. This can be done by editing the information in the pertinent boxes, and then saving as you would a new entry. Content that has been added by the community can be edited. In the image to the left, any of the columns may be edited. Please note that data marked protected has been added by Wiki scripts and can not be edited. However, if you consider something to be wrong or needing to be changed, we encourage you to make your voice heard by editing the table or writing in the section marked Notes below the table.
After entering or editing the information in the table, click on the Save Row button at the bottom of the table.
Any changes you have made are not yet saved at this point.
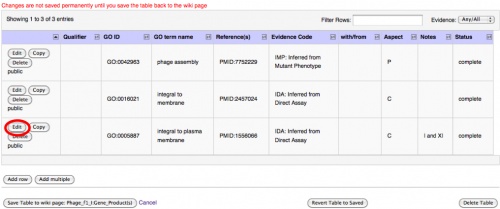
- If you wish to change another row, or go back and edit the annotation you just added, use the indicated Edit button for that row. You can edit information added by yourself or the community, and you can edit a row without adding a row.
- If you wish to add a similar annotation to the same protein (eg, you just added a Component term but also wish to add a Function term from the same paper), you might consider using the Copy button. This will add an exact duplicate, which you can change with the Edit button. You can also use the Add Row and create an entirely new entry for the gene product.
The Edit, Copy and Delete buttons on a row will only execute the given function on that row.
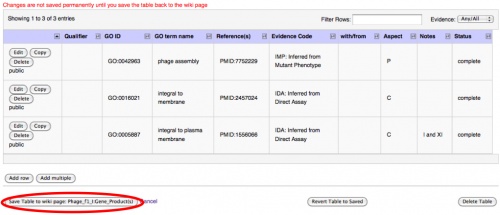
When the row(s) you have added/edited appear to be complete, you need to save again. This is the final save page, where you must save for the second time or your modifications will not be kept. The Save Table to wiki page:__________ button will make your changes appear on the main page; Cancel or Revert Table to Saved will delete your changes. The Cancel button will take you back to the main Gene Products page, whereas the Revert Table to Saved will delete your changes but will keep you on the page where you can edit the table.
Troubleshooting: Editing and Entering Information
- If you would like to make an annotation but are unable to make a "complete" entry with a GO ID, a reference, and an evidence code, consider changing the evidence code. Although the more vague codes (TAS, ND, IC, etc) are discouraged, they signal to the community that an annotation needs closer inspection and likely needs editing. The notes section may be a good place to suggest a more accurate code you can't support with the given evidence, or to indicate that you are currently doing research on the gene and will try to update when possible, etc.
More About the Ontology
The Gene Ontology is actually three ontologies:
- Biological Process (P)
- Cellular Component (C)
- Molecular Function (F)
These three are treated in the same fashion and are only conceptually different. The ontology is structured as an directed acyclic graph, meaning a hierarchal, non-cyclic tree. Sometimes when a term is found to be outside the scope of GO or that could be captured in a better way, that term is then tagged as obsolete; usually there will be recommendations for equivalent usable terms on the GO term page.
More Information
The Gene Ontology website contains much helpful information. A few of the pages we find to be most informative are:
- Structure of the Ontology
- GO Annotation Policies and Guidelines
- Annotation Conventions
- The main Gene Ontology website - http://www.geneontology.org/
- GONUTS - A species-independent wiki for making annotations - http://gowiki.tamu.edu