tsx:On One Page
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;} h2 .editsection { display:none;}</css>
| Quickview | Gene | Gene Product(s) | Expression | Evolution | On One Page |
| Quickview | | Gene | | Product(s) | | Expression| | Evolution | | References | |
Quickview
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
| Standard Name |
tsx |
|---|---|
| Gene Synonym(s) |
ECK0405, b0411, JW0401, T6rec, nupA[1], nupA |
| Product Desc. |
nucleoside channel; receptor of phage T6 and colicin K[2][3] Outer membrane channel for nucleosides; receptor for phage T6 and colicin K[4] |
| Product Synonyms(s) |
nucleoside channel, receptor of phage T6 and colicin K[1], B0411[2][1], NupA[2][1], Tsx[2][1] , ECK0405, JW0401, nupA, T6rec, b0411 |
| Function from GO |
<GO_nr /> |
| Knock-Out Phenotype | |
| Regulation/Expression | |
| Regulation/Activity | |
| Quick Links | |
| edit table |
</protect> See Help:Quickview for help with entering information in the Quickview table. <protect></protect>
Notes
First 22 aa are predicted to be a type I signal peptide. The antibiotic albicidin enters via the Tsx channel.[4]
Gene
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
Nomenclature
See Help:Gene_nomenclature for help with entering information in the Gene Nomenclature table.
| Standard name |
tsx |
|---|---|
| Mnemonic |
T-six |
| Synonyms |
ECK0405, b0411, JW0401, T6rec, nupA[1], nupA |
| edit table |
</protect>
Notes
Location(s) and DNA Sequence
<protect>
See Help:Gene_location for help entering information in the Gene Location and DNA sequence table.
| Strain | Map location | Genome coordinates | Genome browsers | Sequence links |
|---|---|---|---|---|
|
MG1655 |
9.28 minutes |
MG1655: 431237..430353 |
||
|
NC_012967: 400435..399662 |
||||
|
NC_012759: 333112..333996 |
||||
|
W3110 |
|
W3110: 431237..430353 |
||
|
DH10B: 370568..369684 |
||||
| edit table |
</protect>
Notes
Sequence Features
See Help:Gene_sequence_features for help in entering sequence features in EcoliWiki.
| Feature Type | Strain | Genomic Location | Evidence | Reference | Notes |
|---|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Alleles and Phenotypes
See Help:Gene_alleles for how to enter or edit alleles and phenotypes in EcoliWiki.
| Allele | Nt change(s) | AA change(s) | Phenotype: Type | Phenotype: Description | Reference | Availability | Comments |
|---|---|---|---|---|---|---|---|
|
tsx(del) (Keio:JW0401) |
deletion |
deletion |
PMID:16738554 |
||||
|
tsxN276Y |
N276Y |
(in phage resistant mutant) |
Strain variation; seeded from UniProt:P0A927 | ||||
|
tsx-247::Tn10 |
Insertion at 430,667 bp in MG1655 (NC_000913) |
adapted from Nichols et al.[6] |
Synonyms: | ||||
|
tsx-33 |
|||||||
|
tsx-29 |
|||||||
|
tsx-67 |
|||||||
|
tsx-1 |
|||||||
|
tsx-70 |
|||||||
|
tsx-358 |
|||||||
|
tsx-81 |
|||||||
|
tsx-76 |
|||||||
|
tsx-79 |
|||||||
|
tsx-84 |
|||||||
|
tsx-95 |
|||||||
|
tsx-7 |
|||||||
|
tsx-63 |
|||||||
|
tsx-78 |
|||||||
|
tsx-96 |
|||||||
|
tsx-71 |
|||||||
|
tsx-64 |
|||||||
|
tsx-273 |
|||||||
|
tsx-97 |
|||||||
|
tsx-0 |
|||||||
|
tsx-68 |
|||||||
|
tsx-463 |
|||||||
|
tsx-352 |
|||||||
|
tsx-83 |
|||||||
|
tsx-80 |
|||||||
|
tsx-82 |
|||||||
|
tsx-65 |
|||||||
|
tsx-354 |
|||||||
|
tsx-356 |
|||||||
|
tsx-357 |
|||||||
|
tsx-3 |
|||||||
|
tsx-61 |
|||||||
|
tsx-59 |
|||||||
|
tsx-6 |
|||||||
|
tsx-274 |
|||||||
|
tsx-87 |
|||||||
|
tsx-23 |
|||||||
|
tsx-25 |
|||||||
|
tsx-464 |
|||||||
|
tsx-36 |
|||||||
|
tsx-57 |
|||||||
|
tsx-5 |
|||||||
|
tsx-353 |
|||||||
|
tsx-462::Tn10 |
|||||||
|
tsx-93 |
|||||||
|
tsx-466 |
|||||||
|
tsx-69 |
|||||||
|
tsx-465(Am) |
amber (UAG) mutation | ||||||
|
tsx-72 |
|||||||
|
tsx-21 |
|||||||
|
tsx-2 |
|||||||
|
tsx-85 |
|||||||
|
tsx-460 |
|||||||
|
tsx-18 |
|||||||
|
tsx-17 |
|||||||
|
tsx-27 |
|||||||
|
tsx-4 |
|||||||
|
tsx-58 |
|||||||
|
tsx-62 |
|||||||
|
tsx-73 |
|||||||
|
tsx-459 |
|||||||
|
tsx-19 |
|||||||
|
tsx-37 |
|||||||
|
tsx-77 |
|||||||
|
tsx-247::Tn10 |
|||||||
|
tsx-3100::Tn10kan |
|||||||
|
tsx-35 |
|||||||
|
tsx-74 |
|||||||
|
tsx-75 |
|||||||
|
tsx-773(del)::kan |
PMID:16738554 |
||||||
|
tsx mutation in P1807 |
Sensitivity to |
Reduced sensitivity toward albicidin. |
PMID:2191080 |
See Figure 1 B for full experimental results. | |||
|
tsx mutation in strain P1807 |
Metabolic uptake |
Mutation to the tsx region caused a reduced uptake of nucleoside. |
PMID:2191080 |
See figure 2 for full experimental results. | |||
|
tsx mutation in strain GP4 |
Metabolic uptake |
Mutation to the tsx region caused a reduced uptake of ablicidin. |
PMID:2191080 |
These results support the theory that ablicidin resistance is due to reduced uptake of the antibiotic. See figure 3b for experimental results. | |||
|
tsx- in P407 |
Resistant to |
Resistance to Bacteriophage T6 |
PMID:786267 |
Mutation to the tsx gene confers resistance to phage T6, possibly by reducing the production of a protein thought to be involved in phage binding, tsx-protein. | |||
|
tsx--con- in P1731 |
Resistant to |
Resistance to Bacteriophage T6 |
PMID:786267 |
A double mutation to the tsx and con gene confers resistance to phage T6, possibly by reducing the production of a protein thought to be involved in phage binding, tsx-protein. | |||
|
tsx- in P407 |
Resistant to |
Resistance to Colicin K |
PMID:786267 |
Mutation to the tsx gene confers resistance to Colicin K possibly by reducing the production of a specific protein, tsx-protein. | |||
|
tsx--con- in P1731 |
Resistant to |
Resistance to Colicin K |
PMID:786267 |
A double mutation to the tsx and con gene confers resistance to Colicin K, possibly by reducing the production of a protein thought to be involved in phage binding, tsx-protein. | |||
|
tsx-2 in strain KG16 |
Resistant to |
Resistance to strain Phage T6 |
PMID:6808961 |
tsx-2 class of mutants were resistant to bacteriophage T6. This was also seen in strain KG19. | |||
|
tsx-2 in strain KG16 |
Resistant to |
Resistance to Colicin K |
PMID:6808961 |
tsx-2 class of mutants became more resistant to Colicin K. This was also seen in strain KG19. | |||
|
tsx-1 in strain GG24 |
Resistant to |
Resistance to strain Phage T6 |
PMID:6808961 |
tsx-1 class of mutants developed an increased resistance toward bacteriophage T6. This was also seen in strain GG26. | |||
|
tsx-1 in strain GG24d |
Resistant to |
Resistance to Colicin K |
PMID:6808961 |
tsx-2 class of mutants became more resistant to Colicin K. This was also seen in strain GG24. | |||
|
tsx-2 in strain KG16 |
Growth Phenotype |
Muciod growth |
PMID:6808961 |
It was found that the tsx-2 class of mutants produced a Muciod type growth configuration. This included strain KG19. | |||
|
tsx-2 in strain KG16 |
Sensitivity to |
Increased sensitivity to Neomycin. |
PMID:6808961 |
It was found that the tsx-2 class of mutants became increasingly sensitive to aminoglycosides Neomycin and Kanamycin. This included strain KG19. | |||
|
tsx-2 in strain KG16 |
Sensitivity to |
Increased sensitivity to Kanamycin. |
PMID:6808961 |
It was found that the tsx-2 class of mutants became increasingly sensitive to aminoglycosides Neomycin and Kanamycin. This included strain KG19. | |||
|
tsx-2 |
Sensitivity to |
The tsx-2 mutation caused increased sensitivity toward the antibiotic nalidixic acid. |
PMID:6808961 |
Experimental Strain: KG16 |
| ||
| edit table |
<protect></protect>
Notes
Genetic Interactions
<protect>
| Interactor | Interaction | Allele | Score(s) | Reference(s) | Accessions | Notes |
|---|---|---|---|---|---|---|
| edit table |
</protect>
Notes
Genetic Resources
See Help:Gene_resources for help entering information into the Genetic Resources table.
| Resource | Resource Type | Source | Notes/Reference |
|---|---|---|---|
|
JW0401 |
Plasmid clone |
PMID:16769691 Status:Clone OK Primer 1:GCCAAAAAAACATTACTGGCAGC Primer 2:CCGAAGTTGTAACCTACTACCAG | |
|
Kohara Phage |
PMID:3038334 | ||
|
Kohara Phage |
PMID:3038334 | ||
|
Linked marker |
est. P1 cotransduction: 23% [6] | ||
|
Linked marker |
est. P1 cotransduction: 33% [6] | ||
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG11035 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG11035 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120001024 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EchoBASE:EB1028 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0001425 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Product(s)
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Nomenclature
See Help:Product_nomenclature for help entering or editing information in this section of EcoliWiki.
| Standard name |
Tsx |
|---|---|
| Synonyms |
nucleoside channel, receptor of phage T6 and colicin K[1], B0411[2][1], NupA[2][1], Tsx[2][1] , ECK0405, JW0401, nupA, T6rec, b0411 |
| Product description |
nucleoside channel; receptor of phage T6 and colicin K[2][3] Outer membrane channel for nucleosides; receptor for phage T6 and colicin K[4] |
| EC number (for enzymes) |
|
| edit table |
<protect></protect>
Notes
Function
<protect>
Gene Ontology
See Help:Gene_ontology for help entering or editing GO terms and GO annotations in EcoliWiki.
| Qualifier | GO ID | GO term name | Reference | Evidence Code | with/from | Aspect | Notes | Status |
|---|---|---|---|---|---|---|---|---|
|
GO:0005337 |
nucleoside transmembrane transporter activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR003055 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005886 |
plasma membrane |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-1003 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006810 |
transport |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0813 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006811 |
ion transport |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0406 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0009279 |
cell outer membrane |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR003055 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0009279 |
cell outer membrane |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR018013 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0009279 |
cell outer membrane |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0998 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0009597 |
detection of virus |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0580 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0016020 |
membrane |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0472 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0016021 |
integral to membrane |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0812 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0016021 |
integral to membrane |
GO_REF:0000023 |
IEA: Inferred from Electronic Annotation |
SP_SL:SL-9909 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0046718 |
entry of virus into host cell |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0580 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0046930 |
pore complex |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0626 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
| edit table |
Interactions See Help:Product_interactions for help entering or editing information about gene product interactions in this section of EcoliWiki.
| Partner Type | Partner | Notes | References | Evidence |
|---|---|---|---|---|
| edit table |
</protect>
Notes
Localization
See Help:Product_localization for how to add or edit information in this section of EcoliWiki.
| Compartment | Description | Evidence | Reference/Source | Notes | |
|---|---|---|---|---|---|
|
outer membrane |
From EcoCyc[3] |
||||
|
Outer Membrane |
PMID:10806384 |
| |||
| edit table |
<protect></protect>
Notes
Structure and Physical Properties
<protect>
Physical Properties
See Help:Product_physical_properties for help entering or editing information about the physical properties of this gene product.
| Name | |
|---|---|
| Sequence |
MKKTLLAAGA VLALSSSFTV NAAENDKPQY LSDWWHQSVN VVGSYHTRFG PQIRNDTYLE YEAFAKKDWF DFYGYADAPV FFGGNSDAKG IWNHGSPLFM EIEPRFSIDK LTNTDLSFGP FKEWYFANNY IYDMGRNKDG RQSTWYMGLG TDIDTGLPMS LSMNVYAKYQ WQNYGAANEN EWDGYRFKIK YFVPITDLWG GQLSYIGFTN FDWGSDLGDD SGNAINGIKT RTNNSIASSH ILALNYDHWH YSVVARYWHD GGQWNDDAEL NFGNGNFNVR STGWGGYLVV GYNF |
| Length |
294 |
| Mol. Wt |
33.589 kDa |
| pI |
5.0 (calculated) |
| Extinction coefficient |
108,290 (calc based on 21 Y, 14 W, and 0 C residues) |
| edit table |
Domains/Motifs/Modification Sites
|
See Help:Product_domains_motifs for help entering or editing information in this section of EcoliWiki.
|
<motif_map/> |
|
Structure
| </protect>
Structure figures<protect> | ||||||
Notes
Gene Product Resources
See Help:Product_resources for help with entering or editing information in this section of EcoliWiki.
| Resource type | Source | Notes/Reference |
|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
ASAP:ABE-0001425 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EcoCyc:EG11035 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG11035 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
RegulonDB:ECK120001024 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB1028 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Expression
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Overview
This is a placeholder for a summary statement on how expression of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Cellular Levels
| Molecule | Organism or Strain | Value | Units | Experimental Conditions | Assay used | Notes | Reference(s) |
|---|---|---|---|---|---|---|---|
|
Protein |
E. coli K-12 MC4100 |
2.83E+03 |
molecules/cell |
|
emPAI |
PMID:18304323 | |
|
Protein |
E. coli K-12 MG1655 |
14911 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
3927 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
5253 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
| edit table |
Notes
Transcription and Transcriptional Regulation
<protect>
|
See Help:Expression_transcription for help entering or editing information in this section of EcoliWiki.
|
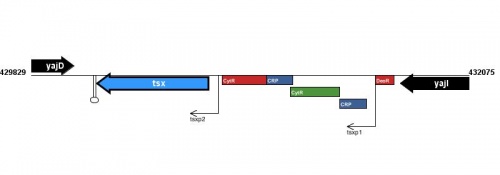 Figure courtesy of RegulonDB |
</protect>
Notes
This is a placeholder for a summary statement about how transcription of this gene is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Translation and Regulation of Translation
<protect><gbrowseImage>
name=NC_000913:431217..431257
source=MG1655
flip=1
type=Gene+DNA_+Protein
preset=Nterminus
</gbrowseImage>
This picture shows the sequence around the N-terminus.
</protect>
Notes
This is a placeholder for a summary statement about how translation of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Turnover and Regulation of Turnover
</protect>
Notes
This is a placeholder for a summary statement about turnover of this gene product. You can help EcoliWiki by becoming a user and writing/editing this statement.
Experimental
<protect>
Mutations Affecting Expression
See Help:Expression_mutations for help entering or editing information in this section of EcoliWiki.
| Allele Name | Mutation | Phenotype | Reference |
|---|---|---|---|
| edit table |
Expression Studies
See Help:Expression_studies for help entering or editing information in this section of EcoliWiki.
| Type | Reference | Notes |
|---|---|---|
|
microarray |
NCBI GEO profiles for tsx | |
|
microarray |
Summary of data for tsx from multiple microarray studies | |
| edit table |
Expression Resources
See Help:Expression_resources for help entering or editing information in this section of EcoliWiki.
| Resource Name | Resource Type | Description | Source | Notes |
|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions Related to tsx Expression
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG11035 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB1028 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG11035 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120001024 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0001425 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Evolution
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Homologs in Other Organisms
See Help:Evolution_homologs for help entering or editing information in this section of EcoliWiki.
| Organism | Homologs (Statistics) | Comments |
|---|---|---|
|
Shigella flexneri |
TSX |
From SHIGELLACYC |
|
E. coli O157 |
TSX |
From ECOO157CYC |
| edit table |
Do-It-Yourself Web Tools
<protect></protect>
Notes
Families
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG11035 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG11035 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120001024 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB1028 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0001425 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
References
See Help:References for how to manage references in EcoliWiki.
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 1.9 Riley, M. et al. (2006) Nucleic Acids Res 34:1-6 (corrected supplemental data from B. Wanner)
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 2.6 2.7 2.8 EcoCyc (release 10.6; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 3.0 3.1 3.2 EcoCyc (release 11.1; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 4.0 4.1 4.2 EcoGene: Rudd, KE (2000) EcoGene: a genome sequence database for Escherichia coli K-12. Nucleic Acids Res 28:60-4.
- ↑ 5.0 5.1 5.2 5.3 CGSC: The Coli Genetics Stock Center
- ↑ 6.0 6.1 6.2 The Tn10 insertion sites determined by Nichols et al. 1998 (PMID:9829956) were reannotated by alignment with E. coli K-12 genome sequence (GenBank accession NC_000913). P1 contransduction frequencies were calculated using the formula of Wu (PMID:5338813).
Categories


