rnr:On One Page
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;} h2 .editsection { display:none;}</css>
| Quickview | Gene | Gene Product(s) | Expression | Evolution | On One Page |
| Quickview | | Gene | | Product(s) | | Expression| | Evolution | | References | |
Quickview
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
| Standard Name |
rnr |
|---|---|
| Gene Synonym(s) |
ECK4175, b4179, JW5741, vacB, yjeC[1], yjeC |
| Product Desc. |
EG11259- Exoribonuclease R; specifically required for SsrA-SmpB-dependent decay of non-stop aberrant mRNAs (Richards, 2006)[2] |
| Product Synonyms(s) |
exoribonuclease R, RNase R[1], B4179[3][1], YjeC[3][1], Rnr[3][1], VacB[3][1] , ECK4175, JW5741, vacB, yjeC, b4179 |
| Function from GO |
<GO_nr /> |
| Knock-Out Phenotype | |
| Regulation/Expression |
transcription unit(s): nsrR-rnr-rlmB[3] |
| Regulation/Activity | |
| Quick Links | |
| edit table |
</protect> See Help:Quickview for help with entering information in the Quickview table. <protect></protect>
Notes
rnr is transiently induced after a cold shock. rnr is orthologous with Shigella virulence gene vacB.[2]
Gene
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
Nomenclature
See Help:Gene_nomenclature for help with entering information in the Gene Nomenclature table.
| Standard name |
rnr |
|---|---|
| Mnemonic |
RNase R |
| Synonyms |
ECK4175, b4179, JW5741, vacB, yjeC[1], yjeC |
| edit table |
</protect>
Notes
Location(s) and DNA Sequence
<protect>
See Help:Gene_location for help entering information in the Gene Location and DNA sequence table.
| Strain | Map location | Genome coordinates | Genome browsers | Sequence links |
|---|---|---|---|---|
|
MG1655 |
94.94 minutes |
MG1655: 4404677..4407118 |
||
|
NC_012967: 4384809..4387250 |
||||
|
NC_012759: 4343412..4345853 |
||||
|
W3110 |
|
W3110: 4411334..4413775 |
||
|
DH10B: 4505039..4507480 |
||||
| edit table |
</protect>
Notes
Sequence Features
See Help:Gene_sequence_features for help in entering sequence features in EcoliWiki.
| Feature Type | Strain | Genomic Location | Evidence | Reference | Notes |
|---|---|---|---|---|---|
|
Coding Start (SO:0000323) |
4404680 |
Edman degradation |
PMID:11948193 |
| |
| edit table |
<protect></protect>
Notes
Alleles and Phenotypes
See Help:Gene_alleles for how to enter or edit alleles and phenotypes in EcoliWiki.
| Allele | Nt change(s) | AA change(s) | Phenotype: Type | Phenotype: Description | Reference | Availability | Comments |
|---|---|---|---|---|---|---|---|
|
Δrnr (Keio:JW5741) |
deletion |
deletion |
PMID:16738554 |
||||
|
Δrnr-729::kan |
PMID:16738554 |
| |||||
| edit table |
<protect></protect>
Notes
Genetic Interactions
<protect>
| Interactor | Interaction | Allele | Score(s) | Reference(s) | Accessions | Notes |
|---|---|---|---|---|---|---|
| edit table |
</protect>
Notes
Genetic Resources
See Help:Gene_resources for help entering information into the Genetic Resources table.
| Resource | Resource Type | Source | Notes/Reference |
|---|---|---|---|
|
JW5741 |
Plasmid clone |
PMID:16769691 Status:Clone OK Primer 1:GCCTCACAAGATCCTTTCCAGGA Primer 2:CCtTCTGCCACTTTTTTCTTCGC | |
|
Kohara Phage |
PMID:3038334 | ||
|
Kohara Phage |
PMID:3038334 | ||
|
Kohara Phage |
PMID:3038334 | ||
|
Linked marker |
est. P1 cotransduction: 11% [5] | ||
|
Linked marker |
est. P1 cotransduction: 45% [5] | ||
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG11259 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG11259 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120001234 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EchoBASE:EB1239 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0013678 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Product(s)
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Nomenclature
See Help:Product_nomenclature for help entering or editing information in this section of EcoliWiki.
| Standard name |
Rnr |
|---|---|
| Synonyms |
exoribonuclease R, RNase R[1], B4179[3][1], YjeC[3][1], Rnr[3][1], VacB[3][1] , ECK4175, JW5741, vacB, yjeC, b4179 |
| Product description |
EG11259- Exoribonuclease R; specifically required for SsrA-SmpB-dependent decay of non-stop aberrant mRNAs (Richards, 2006)[2] |
| EC number (for enzymes) | |
| edit table |
<protect></protect>
Notes
Function
<protect>
Gene Ontology
See Help:Gene_ontology for help entering or editing GO terms and GO annotations in EcoliWiki.
| Qualifier | GO ID | GO term name | Reference | Evidence Code | with/from | Aspect | Notes | Status |
|---|---|---|---|---|---|---|---|---|
|
GO:0005737 |
cytoplasm |
PMID:16397293 |
C |
Seeded from Riley et al 2006 [1]. |
Missing: evidence | |||
|
GO:0003676 |
nucleic acid binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR011129 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003723 |
RNA binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR001900 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003723 |
RNA binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR003029 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003723 |
RNA binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR004476 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003723 |
RNA binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR011805 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003723 |
RNA binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0694 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0004518 |
nuclease activity |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0540 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0004540 |
ribonuclease activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR001900 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0004540 |
ribonuclease activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR004476 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0009405 |
pathogenesis |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0843 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0016070 |
RNA metabolic process |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR004476 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0016787 |
hydrolase activity |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0378 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0004540 |
ribonuclease activity |
PMID:9603904 |
IMP: Inferred from Mutant Phenotype |
F |
Table 2: extracts of rnr mutant cells have reduced poly(A) degrading activity |
complete | ||
|
GO:0016896 |
exoribonuclease activity, producing 5'-phosphomonoesters |
PMID:11948193 |
IDA: Inferred from Direct Assay |
F |
Fig. 3 and Table 2 |
complete | ||
|
GO:0004527 |
exonuclease activity |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0269 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0008997 |
ribonuclease R activity |
PMID:11948193 |
IDA: Inferred from Direct Assay |
F |
Fig. 3 and Table 2 |
complete | ||
| edit table |
Interactions See Help:Product_interactions for help entering or editing information about gene product interactions in this section of EcoliWiki.
| Partner Type | Partner | Notes | References | Evidence |
|---|---|---|---|---|
|
Protein |
cca |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
infC |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
pgm |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
pssA |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
rplA |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rplB |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rplC |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rplF |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
rplI |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rplM |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rplN |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
rplO |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rpsA |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
rpsB |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rpsC |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
rpsD |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rpsE |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rpsF |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rpsG |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rpsM |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
rpsP |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-891136 | |
|
Protein |
intA |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
tgt |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
ycbY |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
yfiF |
PMID:15690043 |
Experiment(s):EBI-883997, EBI-879491 | |
|
Protein |
ygiF |
PMID:15690043 |
Experiment(s):EBI-883997 | |
|
Protein |
ihfA |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
hupA |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rplP |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rplT |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rplU |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rplV |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rplW |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rplX |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rpsH |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rpsI |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rpsK |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rpsN |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rpsO |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rpsR |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rpsT |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rpsU |
PMID:15690043 |
Experiment(s):EBI-891136 | |
|
Protein |
rpsH |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
|
Protein |
rpsP |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):9.921698 | |
|
Protein |
rplP |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
|
Protein |
rpsN |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
|
Protein |
rpsF |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):17.670398 | |
|
Protein |
rpsT |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):2.564045 | |
|
Protein |
hupA |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
|
Protein |
ihfA |
PMID:19402753 |
LCMS(ID Probability):99.0 | |
|
Protein |
rpsM |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):11.502079 | |
|
Protein |
rplU |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
|
Protein |
rplW |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
|
Protein |
rplI |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):30.262063 | |
|
Protein |
rpsI |
PMID:19402753 |
LCMS(ID Probability):99.0 MALDI(Z-score):6.592879 | |
|
Protein |
rpsR |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
|
Protein |
rpsO |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
|
Protein |
rplX |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):3.597110 | |
|
Protein |
rpsK |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):2.388265 | |
|
Protein |
rplB |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):38.617983 | |
|
Protein |
rplM |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):23.975877 | |
|
Protein |
rplO |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):21.852990 | |
|
Protein |
rpsU |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):3.411533 | |
|
Protein |
rpsD |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):35.575045 | |
|
Protein |
rpsC |
PMID:19402753 |
MALDI(Z-score):32.945078 | |
|
Protein |
rplT |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
|
Protein |
rplA |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):50.772162 | |
|
Protein |
infC |
PMID:19402753 |
MALDI(Z-score):20.289543 | |
|
Protein |
rplF |
PMID:19402753 |
LCMS(ID Probability):97.9 MALDI(Z-score):11.134401 | |
|
Protein |
rplD |
PMID:19402753 |
LCMS(ID Probability):99.6 MALDI(Z-score):29.084634 | |
|
Protein |
pssA |
PMID:19402753 |
MALDI(Z-score):19.741475 | |
|
Protein |
yibL |
PMID:19402753 |
LCMS(ID Probability):98.5 | |
|
Protein |
yfiF |
PMID:19402753 |
MALDI(Z-score):26.783167 | |
| edit table |
</protect>
Notes
Localization
See Help:Product_localization for how to add or edit information in this section of EcoliWiki.
| Compartment | Description | Evidence | Reference/Source | Notes |
|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Structure and Physical Properties
<protect>
Physical Properties
See Help:Product_physical_properties for help entering or editing information about the physical properties of this gene product.
| Name | |
|---|---|
| Sequence |
MTKIFIRDDN GGTEMSQDPF QEREAEKYAN PIPSREFILE HLTKREKPAS RDELAVELHI EGEEQLEGLR RRLRAMERDG QLVFTRRQCY ALPERLDLVK GTVIGHRDGY GFLRVEGRKD DLYLSSEQMK TCIHGDQVLA QPLGADRKGR REARIVRVLV PKTSQIVGRY FTEAGVGFVV PDDSRLSFDI LIPPDQIMGA RMGFVVVVEL TQRPTRRTKA VGKIVEVLGD NMGTGMAVDI ALRTHEIPYI WPQAVEQQVA GLKEEVPEEA KAGRVDLRDL PLVTIDGEDA RDFDDAVYCE KKRGGGWRLW VAIADVSYYV RPSTPLDREA RNRGTSVYFP SQVIPMLPEV LSNGLCSLNP QVDRLCMVCE MTVSSKGRLT GYKFYEAVMS SHARLTYTKV WHILQGDQDL REQYAPLVKH LEELHNLYKV LDKAREERGG ISFESEEAKF IFNAERRIER IEQTQRNDAH KLIEECMILA NISAARFVEK AKEPALFRIH DKPSTEAITS FRSVLAELGL ELPGGNKPEP RDYAELLESV ADRPDAEMLQ TMLLRSMKQA IYDPENRGHF GLALQSYAHF TSPIRRYPDL TLHRAIKYLL AKEQGHQGNT TETGGYHYSM EEMLQLGQHC SMAERRADEA TRDVADWLKC DFMLDQVGNV FKGVISSVTG FGFFVRLDDL FIDGLVHVSS LDNDYYRFDQ VGQRLMGESS GQTYRLGDRV EVRVEAVNMD ERKIDFSLIS SERAPRNVGK TAREKAKKGD AGKKGGKRRQ VGKKVNFEPD SAFRGEKKTK PKAAKKDARK AKKPSAKTQK IAAATKAKRA AKKKVAE |
| Length |
827 |
| Mol. Wt |
93.689 kDa |
| pI |
8.8 (calculated) |
| Extinction coefficient |
64,750 - 65,875 (calc based on 25 Y, 5 W, and 9 C residues) |
| edit table |
Domains/Motifs/Modification Sites
|
See Help:Product_domains_motifs for help entering or editing information in this section of EcoliWiki.
|
<motif_map/> |
|
Structure
| </protect>
Structure figures<protect> | ||||||
Notes
Gene Product Resources
See Help:Product_resources for help with entering or editing information in this section of EcoliWiki.
| Resource type | Source | Notes/Reference |
|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
ASAP:ABE-0013678 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EcoCyc:EG11259 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG11259 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
RegulonDB:ECK120001234 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB1239 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Expression
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Overview
This is a placeholder for a summary statement on how expression of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Cellular Levels
| Molecule | Organism or Strain | Value | Units | Experimental Conditions | Assay used | Notes | Reference(s) |
|---|---|---|---|---|---|---|---|
|
Protein |
E. coli K-12 MC4100 |
2.83E+02 |
molecules/cell |
|
emPAI |
PMID:18304323 | |
|
Protein |
Ecoli K-12 |
90.493+/-1.462 |
Molecules/cell |
|
Single Molecule Fluorescence |
PMID:20671182 | |
|
mRNA |
Ecoli K-12 |
0.16048+/-0.01542 |
Molecules/cell |
|
by FISH |
PMID:20671182 | |
|
Protein |
E. coli K-12 MG1655 |
1121 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
605 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
1051 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
| edit table |
Notes
Transcription and Transcriptional Regulation
<protect>
|
See Help:Expression_transcription for help entering or editing information in this section of EcoliWiki.
|
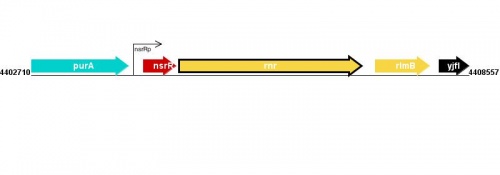 Figure courtesy of RegulonDB |
</protect>
Notes
This is a placeholder for a summary statement about how transcription of this gene is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Translation and Regulation of Translation
<protect><gbrowseImage>
name=NC_000913:4404657..4404697
source=MG1655
type=Gene+DNA_+Protein
preset=Nterminus
</gbrowseImage>
This picture shows the sequence around the N-terminus.
</protect>
Notes
This is a placeholder for a summary statement about how translation of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Turnover and Regulation of Turnover
</protect>
Notes
This is a placeholder for a summary statement about turnover of this gene product. You can help EcoliWiki by becoming a user and writing/editing this statement.
Experimental
<protect>
Mutations Affecting Expression
See Help:Expression_mutations for help entering or editing information in this section of EcoliWiki.
| Allele Name | Mutation | Phenotype | Reference |
|---|---|---|---|
| edit table |
Expression Studies
See Help:Expression_studies for help entering or editing information in this section of EcoliWiki.
| Type | Reference | Notes |
|---|---|---|
|
microarray |
NCBI GEO profiles for rnr | |
|
microarray |
Summary of data for rnr from multiple microarray studies | |
| edit table |
Expression Resources
See Help:Expression_resources for help entering or editing information in this section of EcoliWiki.
| Resource Name | Resource Type | Description | Source | Notes |
|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions Related to rnr Expression
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG11259 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB1239 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG11259 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120001234 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0013678 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Evolution
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Homologs in Other Organisms
See Help:Evolution_homologs for help entering or editing information in this section of EcoliWiki.
| Organism | Homologs (Statistics) | Comments |
|---|---|---|
|
Anopheles gambiae |
|
From Inparanoid:20070104 |
|
Arabidopsis thaliana |
|
From Inparanoid:20070104 |
|
Bos taurus |
|
From Inparanoid:20070104 |
|
Caenorhabditis elegans |
|
From Inparanoid:20070104 |
|
Canis familiaris |
|
From Inparanoid:20070104 |
|
Ciona intestinalis |
|
From Inparanoid:20070104 |
|
Dictyostelium discoideum |
|
From Inparanoid:20070104 |
|
Drosophila melanogaster |
|
From Inparanoid:20070104 |
|
Gallus gallus |
|
From Inparanoid:20070104 |
|
Homo sapiens |
|
From Inparanoid:20070104 |
|
Macaca mulatta |
|
From Inparanoid:20070104 |
|
Monodelphis domestica |
|
From Inparanoid:20070104 |
|
Mus musculus |
|
From Inparanoid:20070104 |
|
Oryza gramene |
|
From Inparanoid:20070104 |
|
Pan troglodytes |
|
From Inparanoid:20070104 |
|
Rattus norvegicus |
|
From Inparanoid:20070104 |
|
Saccharomyces cerevisiae |
|
From Inparanoid:20070104 |
|
Schizosaccharomyces pombe |
|
From Inparanoid:20070104 |
|
Takifugu rubripes |
|
From Inparanoid:20070104 |
|
Tetraodon nigroviridis |
|
From Inparanoid:20070104 |
|
Xenopus tropicalis |
|
From Inparanoid:20070104 |
|
Shigella flexneri |
VACB |
From SHIGELLACYC |
|
E. coli O157 |
VACB |
From ECOO157CYC |
| edit table |
Do-It-Yourself Web Tools
<protect></protect>
Notes
Families
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG11259 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG11259 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120001234 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB1239 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0013678 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
References
See Help:References for how to manage references in EcoliWiki.
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 Riley, M. et al. (2006) Nucleic Acids Res 34:1-6 (corrected supplemental data from B. Wanner)
- ↑ 2.0 2.1 2.2 EcoGene: Rudd, KE (2000) EcoGene: a genome sequence database for Escherichia coli K-12. Nucleic Acids Res 28:60-4.
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 3.6 3.7 3.8 EcoCyc (release 10.6; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 4.0 4.1 4.2 CGSC: The Coli Genetics Stock Center
- ↑ 5.0 5.1 The Tn10 insertion sites determined by Nichols et al. 1998 (PMID:9829956) were reannotated by alignment with E. coli K-12 genome sequence (GenBank accession NC_000913). P1 contransduction frequencies were calculated using the formula of Wu (PMID:5338813).
Categories
- Genes with homologs in Anopheles gambiae
- Genes with homologs in Arabidopsis thaliana
- Genes with homologs in Bos taurus
- Genes with homologs in Caenorhabditis elegans
- Genes with homologs in Canis familiaris
- Genes with homologs in Ciona intestinalis
- Genes with homologs in Dictyostelium discoideum
- Genes with homologs in Drosophila melanogaster
- Genes with homologs in Gallus gallus
- Genes with homologs in Homo sapiens
- Genes with homologs in Macaca mulatta
- Genes with homologs in Monodelphis domestica
- Genes with homologs in Mus musculus
- Genes with homologs in Oryza gramene
- Genes with homologs in Pan troglodytes
- Genes with homologs in Rattus norvegicus
- Genes with homologs in Saccharomyces cerevisiae
- Genes with homologs in Schizosaccharomyces pombe
- Genes with homologs in Takifugu rubripes
- Genes with homologs in Tetraodon nigroviridis
- Genes with homologs in Xenopus tropicalis
- Genes with homologs in Shigella flexneri
- Genes with homologs in E. coli O157


