malK:On One Page
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;} h2 .editsection { display:none;}</css>
| Quickview | Gene | Gene Product(s) | Expression | Evolution | On One Page |
| Quickview | | Gene | | Product(s) | | Expression| | Evolution | | References | |
Quickview
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
| Standard Name |
malK |
|---|---|
| Gene Synonym(s) |
ECK4027, b4035, JW3995, malB[1] |
| Product Desc. |
Component of maltose ABC transporter[2][3]; Maltose/Maltodextrin Transport System[3] Maltose transport complex, ATP-binding subunit[4] |
| Product Synonyms(s) |
fused maltose transport subunit, ATP-binding component of ABC superfamily[1], regulatory protein[1], B4035[2][1], MalB[2][1], MalK[2][1] , ECK4027, JW3995, b4035 |
| Function from GO |
<GO_nr /> |
| Knock-Out Phenotype | |
| Regulation/Expression |
transcription unit(s): malK-lamB-malM[2], OP00165 |
| Regulation/Activity | |
| Quick Links | |
| edit table |
</protect> See Help:Quickview for help with entering information in the Quickview table. <protect></protect>
Notes
The inactive, ATP-bound form of MalK directly antagonizes the mal regulon activator MalT. The maltose transport complex MalFGK(2) also transports maltodextrin. Overexpression causes filamentous biofilm formation. MalT regulon.[4]
Gene
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
Nomenclature
See Help:Gene_nomenclature for help with entering information in the Gene Nomenclature table.
| Standard name |
malK |
|---|---|
| Mnemonic |
Maltose |
| Synonyms |
ECK4027, b4035, JW3995, malB[1] |
| edit table |
</protect>
Notes
Location(s) and DNA Sequence
<protect>
See Help:Gene_location for help entering information in the Gene Location and DNA sequence table.
| Strain | Map location | Genome coordinates | Genome browsers | Sequence links |
|---|---|---|---|---|
|
MG1655 |
91.49 minutes |
MG1655: 4244807..4245922 |
||
|
NC_012967: 4225716..4226831 |
||||
|
NC_012759: 4183652..4184767 |
||||
|
W3110 |
|
W3110: 4250374..4251489 |
||
|
DH10B: 4344503..4345618 |
||||
| edit table |
</protect>
Notes
Sequence Features
See Help:Gene_sequence_features for help in entering sequence features in EcoliWiki.
| Feature Type | Strain | Genomic Location | Evidence | Reference | Notes |
|---|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Alleles and Phenotypes
See Help:Gene_alleles for how to enter or edit alleles and phenotypes in EcoliWiki.
| Allele | Nt change(s) | AA change(s) | Phenotype: Type | Phenotype: Description | Reference | Availability | Comments |
|---|---|---|---|---|---|---|---|
|
ΔmalK (Keio:JW3995) |
deletion |
deletion |
PMID:16738554 |
||||
|
malKG137A |
G137A |
Loss of maltose transport. Has greater ability to decrease mal gene expression than wild-type malK |
seeded from UniProt:P68187 | ||||
|
malKA124T |
A124T |
Resistant to inhibitory effects of alpha-methylglucoside but retains transport capacity |
seeded from UniProt:P68187 | ||||
|
malKA85M |
A85M |
Suppressor of EAA loop mutations in MalFG |
seeded from UniProt:P68187 | ||||
|
malKK106C |
K106C |
Suppressor of EAA loop mutations in MalFG |
seeded from UniProt:P68187 | ||||
|
malKV114C |
V114C |
Suppressor of EAA loop mutations in MalFG |
seeded from UniProt:P68187 | ||||
|
malKR228C |
R228C |
Resistant to inhibitory effects of alpha-methylglucoside but retains transport capacity |
seeded from UniProt:P68187 | ||||
|
malKF241I |
F241I |
Resistant to inhibitory effects of alpha-methylglucoside but retains transport capacity |
seeded from UniProt:P68187 | ||||
|
malKW267G |
W267G |
Normal maltose transport but constitutive mal gene expression |
seeded from UniProt:P68187 | ||||
|
malKD158N |
D158N |
Loss of maltose transport but retains ability to repress mal genes |
seeded from UniProt:P68187 | ||||
|
malKG340A |
G340A |
Maltose transport is affected but retains ability to interact with malT |
seeded from UniProt:P68187 | ||||
|
malKG278P |
G278P |
Resistant to inhibitory effects of alpha-methylglucoside but retains transport capacity |
seeded from UniProt:P68187 | ||||
|
malKS282L |
S282L |
Resistant to inhibitory effects of alpha-methylglucoside but retains transport capacity |
seeded from UniProt:P68187 | ||||
|
malKG284S |
G284S |
Resistant to inhibitory effects of alpha-methylglucoside but retains transport capacity |
seeded from UniProt:P68187 | ||||
|
malKG302D |
G302D |
Resistant to inhibitory effects of alpha-methylglucoside but retains transport capacity |
seeded from UniProt:P68187 | ||||
|
malKE308Q |
E308Q |
Maltose transport is affected but retains ability to interact with malT |
seeded from UniProt:P68187 | ||||
|
malKS322F |
S322F |
Resistant to inhibitory effects of alpha-methylglucoside but retains transport capacity |
seeded from UniProt:P68187 | ||||
|
malKG346S |
G346S |
Normal maltose transport but constitutive mal gene expression |
seeded from UniProt:P68187 | ||||
|
malKF355Y |
F355Y |
Maltose transport is affected but retains ability to interact with malT |
seeded from UniProt:P68187 | ||||
|
malKV117M |
V117M |
Suppressor of EAA loop mutations in MalFG |
seeded from UniProt:P68187 | ||||
|
malKE119K |
E119K |
Resistant to inhibitory effects of alpha-methylglucoside but retains transport capacity |
seeded from UniProt:P68187 | ||||
|
malK62(Am) |
amber (UAG) mutation | ||||||
|
malK58::Tn5 |
|||||||
|
malK27 |
|||||||
|
ΔmalK731::kan |
PMID:16738554 |
| |||||
| edit table |
<protect></protect>
Notes
Genetic Interactions
<protect>
| Interactor | Interaction | Allele | Score(s) | Reference(s) | Accessions | Notes |
|---|---|---|---|---|---|---|
| edit table |
</protect>
Notes
Genetic Resources
See Help:Gene_resources for help entering information into the Genetic Resources table.
| Resource | Resource Type | Source | Notes/Reference |
|---|---|---|---|
|
JW3995 |
Plasmid clone |
PMID:16769691 Status:Clone OK Primer 1:GCCGCGAGCGTACAGCTGCAAAA Primer 2:CCAACGCCCGGCTCCTTATGCAG | |
|
Kohara Phage |
PMID:3038334 | ||
|
Kohara Phage |
PMID:3038334 | ||
|
Kohara Phage |
PMID:3038334 | ||
|
Linked marker |
est. P1 cotransduction: 87% [6] | ||
|
Linked marker |
est. P1 cotransduction: 4% [6] | ||
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG10558 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10558 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120000551 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EchoBASE:EB0553 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0013216 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Product(s)
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Nomenclature
See Help:Product_nomenclature for help entering or editing information in this section of EcoliWiki.
| Standard name |
MalK |
|---|---|
| Synonyms |
fused maltose transport subunit, ATP-binding component of ABC superfamily[1], regulatory protein[1], B4035[2][1], MalB[2][1], MalK[2][1] , ECK4027, JW3995, b4035 |
| Product description |
Component of maltose ABC transporter[2][3]; Maltose/Maltodextrin Transport System[3] Maltose transport complex, ATP-binding subunit[4] |
| EC number (for enzymes) |
|
| edit table |
<protect></protect>
Notes
Function
<protect>
Gene Ontology
See Help:Gene_ontology for help entering or editing GO terms and GO annotations in EcoliWiki.
| Qualifier | GO ID | GO term name | Reference | Evidence Code | with/from | Aspect | Notes | Status |
|---|---|---|---|---|---|---|---|---|
|
GO:0000166 |
nucleotide binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR003593 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0000166 |
nucleotide binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0547 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR003439 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR008995 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR013611 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR015855 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR017871 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0067 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005886 |
plasma membrane |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0997 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005886 |
plasma membrane |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-1003 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0015423 |
maltose-transporting ATPase activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR015855 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0015423 |
maltose-transporting ATPase activity |
GOA:spec |
IEA: Inferred from Electronic Annotation |
EC:3.6.3.19 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0015609 |
maltooligosaccharide-importing ATPase activity |
GOA:hamap |
IEA: Inferred from Electronic Annotation |
HAMAP:MF_01709 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0015768 |
maltose transport |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR015855 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0019898 |
extrinsic to membrane |
GO_REF:0000023 |
IEA: Inferred from Electronic Annotation |
SP_SL:SL-9903 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0043190 |
ATP-binding cassette (ABC) transporter complex |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR008995 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0043190 |
ATP-binding cassette (ABC) transporter complex |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR013611 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
| edit table |
Interactions See Help:Product_interactions for help entering or editing information about gene product interactions in this section of EcoliWiki.
| Partner Type | Partner | Notes | References | Evidence |
|---|---|---|---|---|
|
Protein |
Subunits of maltose ABC transporter |
could be indirect |
||
|
Protein |
Subunits of Maltose/Maltodextrin Transport System |
could be indirect |
| |
| edit table |
</protect>
Notes
Localization
See Help:Product_localization for how to add or edit information in this section of EcoliWiki.
| Compartment | Description | Evidence | Reference/Source | Notes |
|---|---|---|---|---|
|
cytoplasm |
From EcoCyc[3] |
| ||
| edit table |
<protect></protect>
Notes
Structure and Physical Properties
<protect>
Physical Properties
See Help:Product_physical_properties for help entering or editing information about the physical properties of this gene product.
| Name | |
|---|---|
| Sequence |
MASVQLQNVT KAWGEVVVSK DINLDIHEGE FVVFVGPSGC GKSTLLRMIA GLETITSGDL FIGEKRMNDT PPAERGVGMV FQSYALYPHL SVAENMSFGL KLAGAKKEVI NQRVNQVAEV LQLAHLLDRK PKALSGGQRQ RVAIGRTLVA EPSVFLLDEP LSNLDAALRV QMRIEISRLH KRLGRTMIYV THDQVEAMTL ADKIVVLDAG RVAQVGKPLE LYHYPADRFV AGFIGSPKMN FLPVKVTATA IDQVQVELPM PNRQQVWLPV ESRDVQVGAN MSLGIRPEHL LPSDIADVIL EGEVQVVEQL GNETQIHIQI PSIRQNLVYR QNDVVLVEEG ATFAIGLPPE RCHLFREDGT ACRRLHKEPG V |
| Length |
371 |
| Mol. Wt |
40.989 kDa |
| pI |
6.7 (calculated) |
| Extinction coefficient |
19,940 - 20,315 (calc based on 6 Y, 2 W, and 3 C residues) |
| edit table |
Domains/Motifs/Modification Sites
|
See Help:Product_domains_motifs for help entering or editing information in this section of EcoliWiki.
|
<motif_map/> |
|
Structure
| </protect>
Structure figures<protect> | ||||||
Notes
Gene Product Resources
See Help:Product_resources for help with entering or editing information in this section of EcoliWiki.
| Resource type | Source | Notes/Reference |
|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
ASAP:ABE-0013216 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EcoCyc:EG10558 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10558 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
RegulonDB:ECK120000551 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB0553 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Expression
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Overview
This is a placeholder for a summary statement on how expression of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Cellular Levels
| Molecule | Organism or Strain | Value | Units | Experimental Conditions | Assay used | Notes | Reference(s) |
|---|---|---|---|---|---|---|---|
|
Protein |
E. coli K-12 MC4100 |
2.12E+02 |
molecules/cell |
|
emPAI |
PMID:18304323 | |
|
Protein |
Ecoli K-12 |
96.891+/-0.434 |
Molecules/cell |
|
Single Molecule Fluorescence |
PMID:20671182 | |
|
mRNA |
Ecoli K-12 |
0.70273+/-0.09491 |
Molecules/cell |
|
by FISH |
PMID:20671182 | |
|
mRNA |
Ecoli K-12 |
0.059641256 |
Molecules/cell |
|
by RNA_Seq |
PMID:20671182 | |
|
Protein |
E. coli K-12 MG1655 |
31 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
61 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
0a |
molecules/cell/generation |
|
Ribosome Profiling |
Low confidence in the sequencing data set. |
PMID: 24766808 |
| edit table |
Notes
Transcription and Transcriptional Regulation
<protect>
|
See Help:Expression_transcription for help entering or editing information in this section of EcoliWiki.
|
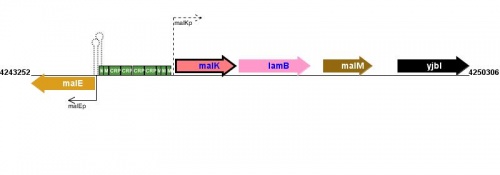 Figure courtesy of RegulonDB |
</protect>
Notes
This is a placeholder for a summary statement about how transcription of this gene is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Translation and Regulation of Translation
<protect><gbrowseImage>
name=NC_000913:4244787..4244827
source=MG1655
type=Gene+DNA_+Protein
preset=Nterminus
</gbrowseImage>
This picture shows the sequence around the N-terminus.
</protect>
Notes
This is a placeholder for a summary statement about how translation of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Turnover and Regulation of Turnover
</protect>
Notes
This is a placeholder for a summary statement about turnover of this gene product. You can help EcoliWiki by becoming a user and writing/editing this statement.
Experimental
<protect>
Mutations Affecting Expression
See Help:Expression_mutations for help entering or editing information in this section of EcoliWiki.
| Allele Name | Mutation | Phenotype | Reference |
|---|---|---|---|
| edit table |
Expression Studies
See Help:Expression_studies for help entering or editing information in this section of EcoliWiki.
| Type | Reference | Notes |
|---|---|---|
|
microarray |
NCBI GEO profiles for malK | |
|
microarray |
Summary of data for malK from multiple microarray studies | |
| edit table |
Expression Resources
See Help:Expression_resources for help entering or editing information in this section of EcoliWiki.
| Resource Name | Resource Type | Description | Source | Notes |
|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions Related to malK Expression
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG10558 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB0553 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10558 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120000551 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0013216 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Evolution
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Homologs in Other Organisms
See Help:Evolution_homologs for help entering or editing information in this section of EcoliWiki.
| Organism | Homologs (Statistics) | Comments |
|---|---|---|
|
Anopheles gambiae |
|
From Inparanoid:20070104 |
|
Drosophila melanogaster |
|
From Inparanoid:20070104 |
|
Drosophila pseudoobscura |
|
From Inparanoid:20070104 |
|
Takifugu rubripes |
|
From Inparanoid:20070104 |
|
Tetraodon nigroviridis |
|
From Inparanoid:20070104 |
|
Xenopus tropicalis |
|
From Inparanoid:20070104 |
|
Shigella flexneri |
MALK |
From SHIGELLACYC |
|
E. coli O157 |
MALK |
From ECOO157CYC |
| edit table |
Do-It-Yourself Web Tools
<protect></protect>
Notes
Families
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG10558 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10558 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120000551 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB0553 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0013216 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
References
See Help:References for how to manage references in EcoliWiki.
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 Riley, M. et al. (2006) Nucleic Acids Res 34:1-6 (corrected supplemental data from B. Wanner)
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 EcoCyc (release 10.6; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 3.6 EcoCyc (release 11.1; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 4.0 4.1 4.2 EcoGene: Rudd, KE (2000) EcoGene: a genome sequence database for Escherichia coli K-12. Nucleic Acids Res 28:60-4.
- ↑ 5.0 5.1 5.2 CGSC: The Coli Genetics Stock Center
- ↑ 6.0 6.1 The Tn10 insertion sites determined by Nichols et al. 1998 (PMID:9829956) were reannotated by alignment with E. coli K-12 genome sequence (GenBank accession NC_000913). P1 contransduction frequencies were calculated using the formula of Wu (PMID:5338813).
Categories
- Genes with homologs in Anopheles gambiae
- Genes with homologs in Drosophila melanogaster
- Genes with homologs in Drosophila pseudoobscura
- Genes with homologs in Takifugu rubripes
- Genes with homologs in Tetraodon nigroviridis
- Genes with homologs in Xenopus tropicalis
- Genes with homologs in Shigella flexneri
- Genes with homologs in E. coli O157


