ada:On One Page
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;} h2 .editsection { display:none;}</css>
| Quickview | Gene | Gene Product(s) | Expression | Evolution | On One Page |
| Quickview | | Gene | | Product(s) | | Expression| | Evolution | | References | |
Quickview
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
| Standard Name |
ada |
|---|---|
| Gene Synonym(s) |
ECK2205, b2213, JW2201[1], JW2201 |
| Product Desc. |
Ada transcriptional dual regulator[2][3] O6-methylguanine-DNA methyltransferase, transcription factor; inducible; DNA repair against methylating and alkylating agents; binds Zn(II)[4] |
| Product Synonyms(s) |
fused DNA-binding transcriptional dual regulator[1], O6-methylguanine-DNA methyltransferase[1], B2213[2][1], Ada[2][1] , ECK2205, JW2201, b2213 |
| Function from GO |
<GO_nr /> |
| Knock-Out Phenotype | |
| Regulation/Expression | |
| Regulation/Activity | |
| Quick Links | |
| edit table |
</protect> See Help:Quickview for help with entering information in the Quickview table. <protect></protect>
Notes
The Ada N-terminal domain acts as a transcriptional activator of ada, alkA, alkB and aidB. Ogt, Ada-C and YbaZ are homologous.[4]
Gene
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
Nomenclature
See Help:Gene_nomenclature for help with entering information in the Gene Nomenclature table.
| Standard name |
ada |
|---|---|
| Mnemonic |
Adaptive (response) |
| Synonyms |
ECK2205, b2213, JW2201[1], JW2201 |
| edit table |
</protect>
Notes
Location(s) and DNA Sequence
<protect>
See Help:Gene_location for help entering information in the Gene Location and DNA sequence table.
| Strain | Map location | Genome coordinates | Genome browsers | Sequence links |
|---|---|---|---|---|
|
MG1655 |
49.73 minutes |
MG1655: 2308427..2307363 |
||
|
NC_012967: 2260509..2259445 |
||||
|
NC_012759: 2193168..2194232 |
||||
|
W3110 |
|
W3110: 2313739..2312675 |
||
|
DH10B: 2399415..2398351 |
||||
| edit table |
</protect>
Notes
Sequence Features
See Help:Gene_sequence_features for help in entering sequence features in EcoliWiki.
| Feature Type | Strain | Genomic Location | Evidence | Reference | Notes |
|---|---|---|---|---|---|
|
Coding Start (SO:0000323) |
2307363 |
Edman degradation |
PMID:2987251 |
| |
| edit table |
<protect></protect>
Notes
Alleles and Phenotypes
See Help:Gene_alleles for how to enter or edit alleles and phenotypes in EcoliWiki.
| Allele | Nt change(s) | AA change(s) | Phenotype: Type | Phenotype: Description | Reference | Availability | Comments |
|---|---|---|---|---|---|---|---|
|
adaE75D |
E75D |
(in strain: B) |
Strain variation; seeded from UniProt:P06134 | ||||
|
adaAQ79PR |
AQ79PR |
(in strain: B) |
Strain variation; seeded from UniProt:P06134 | ||||
|
adaI318V |
I318V |
(in strain: B) |
Strain variation; seeded from UniProt:P06134 | ||||
|
adaT330S |
T330S |
(in strain: B) |
Strain variation; seeded from UniProt:P06134 | ||||
|
Δada (Keio:JW2201) |
deletion |
deletion |
PMID:16738554 |
||||
|
ada::Tn5KAN-2 (FB20708) |
Insertion at nt 687 in Minus orientation |
PMID:15262929 |
does not contain pKD46 | ||||
|
ada::Tn5KAN-2 (FB20709) |
Insertion at nt 687 in Minus orientation |
PMID:15262929 |
contains pKD46 | ||||
|
ada-1 |
|||||||
|
ada-2 |
|||||||
|
ada-3 |
|||||||
|
ada-4 |
|||||||
|
ada-5 |
|||||||
|
ada-6 |
|||||||
|
ada-766(del)::kan |
PMID:16738554 |
| |||||
| edit table |
<protect></protect>
Notes
Genetic Interactions
<protect>
| Interactor | Interaction | Allele | Score(s) | Reference(s) | Accessions | Notes |
|---|---|---|---|---|---|---|
| edit table |
</protect>
Notes
Genetic Resources
See Help:Gene_resources for help entering information into the Genetic Resources table.
| Resource | Resource Type | Source | Notes/Reference |
|---|---|---|---|
|
JW2201 |
Plasmid clone |
PMID:16769691 Status:Clone OK Primer 1:GCCAAAAAAGCCACATGCTTAAC Primer 2:CCtCTCTCCTCATTTTCAGCTTC | |
|
Kohara Phage |
PMID:3038334 | ||
|
Linked marker |
est. P1 cotransduction: 82% [6] | ||
|
Linked marker |
est. P1 cotransduction: 65% [6] | ||
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG10029 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10029 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120000026 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EchoBASE:EB0028 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0007311 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Product(s)
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Nomenclature
See Help:Product_nomenclature for help entering or editing information in this section of EcoliWiki.
| Standard name |
Ada |
|---|---|
| Synonyms |
fused DNA-binding transcriptional dual regulator[1], O6-methylguanine-DNA methyltransferase[1], B2213[2][1], Ada[2][1] , ECK2205, JW2201, b2213 |
| Product description |
Ada transcriptional dual regulator[2][3] O6-methylguanine-DNA methyltransferase, transcription factor; inducible; DNA repair against methylating and alkylating agents; binds Zn(II)[4] |
| EC number (for enzymes) | |
| edit table |
<protect></protect>
Notes
Function
<protect>
Gene Ontology
See Help:Gene_ontology for help entering or editing GO terms and GO annotations in EcoliWiki.
| Qualifier | GO ID | GO term name | Reference | Evidence Code | with/from | Aspect | Notes | Status |
|---|---|---|---|---|---|---|---|---|
|
GO:0003677 |
DNA binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR004026 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003677 |
DNA binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0238 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003700 |
transcription factor activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR000005 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003824 |
catalytic activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR014048 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003908 |
methylated-DNA-(protein)-cysteine S-methyltransferase activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR001497 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003908 |
methylated-DNA-(protein)-cysteine S-methyltransferase activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR008332 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003908 |
methylated-DNA-(protein)-cysteine S-methyltransferase activity |
GOA:spec |
IEA: Inferred from Electronic Annotation |
EC:2.1.1.63 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005622 |
intracellular |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR000005 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006281 |
DNA repair |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR001497 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006281 |
DNA repair |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR004026 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006281 |
DNA repair |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR008332 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006281 |
DNA repair |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR014048 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006281 |
DNA repair |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0234 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006350 |
transcription |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0804 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006355 |
regulation of transcription, DNA-dependent |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR000005 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006355 |
regulation of transcription, DNA-dependent |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR004026 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006974 |
response to DNA damage stimulus |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0227 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0008270 |
zinc ion binding |
PMID:1581309 |
IDA: Inferred from Direct Assay |
F |
N-terminal domain binds Zn |
complete | ||
|
GO:0008168 |
methyltransferase activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR004026 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0008168 |
methyltransferase activity |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0489 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0008270 |
zinc ion binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR004026 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0008270 |
zinc ion binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0862 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0008270 |
zinc ion binding |
PMID:1581309 |
IDA: Inferred from Direct Assay |
F |
Seeded from EcoCyc (v14.0) |
complete | ||
|
GO:0016740 |
transferase activity |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0808 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0043565 |
sequence-specific DNA binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR000005 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0003908 |
methylated-DNA-(protein)-cysteine S-methyltransferase activity |
PMID:6754717 |
IDA: Inferred from Direct Assay |
F |
complete | |||
|
GO:0005625 |
soluble fraction |
PMID:6754717 |
IDA: Inferred from Direct Assay |
C |
complete | |||
|
GO:0006307 |
DNA dealkylation involved in DNA repair |
PMID:6754717 |
IDA: Inferred from Direct Assay |
P |
complete | |||
|
GO:0006307 |
DNA dealkylation involved in DNA repair |
PMID: 282622 |
IDA: Inferred from Direct Assay |
P |
complete | |||
Interactions See Help:Product_interactions for help entering or editing information about gene product interactions in this section of EcoliWiki.
| Partner Type | Partner | Notes | References | Evidence |
|---|---|---|---|---|
|
Protein |
rplB |
PMID:16606699 |
Experiment(s):EBI-1142195 | |
|
Protein |
rpsC |
PMID:16606699 |
Experiment(s):EBI-1142195 | |
|
Protein |
moeB |
PMID:16606699 |
Experiment(s):EBI-1142195 | |
|
Protein |
fur |
PMID:16606699 |
Experiment(s):EBI-1142195 | |
|
Protein |
rpsG |
PMID:16606699 |
Experiment(s):EBI-1142195 | |
|
Protein |
ybiA |
PMID:16606699 |
Experiment(s):EBI-1142195 | |
| edit table |
</protect>
Notes
Localization
See Help:Product_localization for how to add or edit information in this section of EcoliWiki.
| Compartment | Description | Evidence | Reference/Source | Notes |
|---|---|---|---|---|
|
membrane |
From EcoCyc[3] |
| ||
| edit table |
<protect></protect>
Notes
Structure and Physical Properties
<protect>
Physical Properties
See Help:Product_physical_properties for help entering or editing information about the physical properties of this gene product.
| Name | |
|---|---|
| Sequence |
MKKATCLTDD QRWQSVLARD PNADGEFVFA VRTTGIFCRP SCRARHALRE NVSFYANASE ALAAGFRPCK RCQPEKANAQ QHRLDKITHA CRLLEQETPV TLEALADQVA MSPFHLHRLF KATTGMTPKA WQQAWRARRL RESLAKGESV TTSILNAGFP DSSSYYRKAD ETLGMTAKQF RHGGENLAVR YALADCELGR CLVAESERGI CAILLGDDDA TLISELQQMF PAADNAPADL MFQQHVREVI ASLNQRDTPL TLPLDIRGTA FQQQVWQALR TIPCGETVSY QQLANAIGKP KAVRAVASAC AANKLAIIIP CHRVVRGDGT LSGYRWGVSR KAQLLRREAE NEER |
| Length |
354 |
| Mol. Wt |
39.323 kDa |
| pI |
8.8 (calculated) |
| Extinction coefficient |
36,440 - 37,940 (calc based on 6 Y, 5 W, and 12 C residues) |
| edit table |
Domains/Motifs/Modification Sites
|
See Help:Product_domains_motifs for help entering or editing information in this section of EcoliWiki.
|
<motif_map/> |
|
Structure
| </protect>
Structure figures<protect> | ||||||
Notes
Gene Product Resources
See Help:Product_resources for help with entering or editing information in this section of EcoliWiki.
| Resource type | Source | Notes/Reference |
|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
ASAP:ABE-0007311 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EcoCyc:EG10029 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10029 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
RegulonDB:ECK120000026 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB0028 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Expression
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Overview
This is a placeholder for a summary statement on how expression of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Cellular Levels
| Molecule | Organism or Strain | Value | Units | Experimental Conditions | Assay used | Notes | Reference(s) |
|---|---|---|---|---|---|---|---|
|
Protein |
E. coli K-12 MG1655 |
21 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
15 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
8a |
molecules/cell/generation |
|
Ribosome Profiling |
Low confidence in the sequencing data set. |
PMID: 24766808 |
| edit table |
Notes
Transcription and Transcriptional Regulation
<protect>
|
See Help:Expression_transcription for help entering or editing information in this section of EcoliWiki.
|
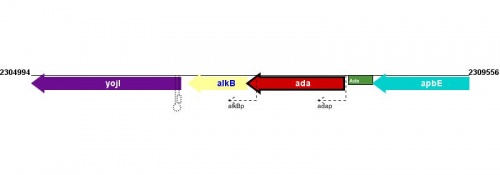 Figure courtesy of RegulonDB |
</protect>
Notes
This is a placeholder for a summary statement about how transcription of this gene is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Translation and Regulation of Translation
<protect><gbrowseImage>
name=NC_000913:2308407..2308447
source=MG1655
flip=1
type=Gene+DNA_+Protein
preset=Nterminus
</gbrowseImage>
This picture shows the sequence around the N-terminus.
</protect>
Notes
This is a placeholder for a summary statement about how translation of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Turnover and Regulation of Turnover
</protect>
Notes
This is a placeholder for a summary statement about turnover of this gene product. You can help EcoliWiki by becoming a user and writing/editing this statement.
Experimental
<protect>
Mutations Affecting Expression
See Help:Expression_mutations for help entering or editing information in this section of EcoliWiki.
| Allele Name | Mutation | Phenotype | Reference |
|---|---|---|---|
| edit table |
Expression Studies
See Help:Expression_studies for help entering or editing information in this section of EcoliWiki.
| Type | Reference | Notes |
|---|---|---|
|
microarray |
NCBI GEO profiles for ada | |
|
microarray |
Summary of data for ada from multiple microarray studies | |
| edit table |
Expression Resources
See Help:Expression_resources for help entering or editing information in this section of EcoliWiki.
| Resource Name | Resource Type | Description | Source | Notes |
|---|---|---|---|---|
|
GFP Fusion |
Intergenic region (2308332..2308580) fused to gfpmut2. |
GFP fusion described in Zaslaver, et al. | ||
| edit table |
<protect></protect>
Notes
Accessions Related to ada Expression
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:EG10029 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB0028 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10029 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120000026 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0007311 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Evolution
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Homologs in Other Organisms
See Help:Evolution_homologs for help entering or editing information in this section of EcoliWiki.
| Organism | Homologs (Statistics) | Comments |
|---|---|---|
|
Apis mellifera |
|
From Inparanoid:20070104 |
|
Caenorhabditis briggsae |
|
From Inparanoid:20070104 |
|
Caenorhabditis elegans |
|
From Inparanoid:20070104 |
|
Canis familiaris |
|
From Inparanoid:20070104 |
|
Dictyostelium discoideum |
|
From Inparanoid:20070104 |
|
Drosophila melanogaster |
|
From Inparanoid:20070104 |
|
Drosophila pseudoobscura |
|
From Inparanoid:20070104 |
|
Homo sapiens |
|
From Inparanoid:20070104 |
|
Macaca mulatta |
|
From Inparanoid:20070104 |
|
Mus musculus |
|
From Inparanoid:20070104 |
|
Pan troglodytes |
|
From Inparanoid:20070104 |
|
Rattus norvegicus |
|
From Inparanoid:20070104 |
|
Takifugu rubripes |
|
From Inparanoid:20070104 |
|
Tetraodon nigroviridis |
|
From Inparanoid:20070104 |
|
Shigella flexneri |
ADA |
From SHIGELLACYC |
|
E. coli O157 |
ADA |
From ECOO157CYC |
| edit table |
Do-It-Yourself Web Tools
<protect></protect>
Notes
Families
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
PF00165 Bacterial regulatory helix-turn-helix proteins, AraC family |
||
|
PF00165 Bacterial regulatory helix-turn-helix proteins, AraC family |
||
|
PF02870 6-O-methylguanine DNA methyltransferase, ribonuclease-like domain |
||
|
PF01035 6-O-methylguanine DNA methyltransferase, DNA binding domain |
||
|
EcoCyc:EG10029 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG10029 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120000026 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB0028 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0007311 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
References
See Help:References for how to manage references in EcoliWiki.
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 Riley, M. et al. (2006) Nucleic Acids Res 34:1-6 (corrected supplemental data from B. Wanner)
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 2.6 EcoCyc (release 10.6; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 3.0 3.1 3.2 EcoCyc (release 11.1; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 4.0 4.1 4.2 EcoGene: Rudd, KE (2000) EcoGene: a genome sequence database for Escherichia coli K-12. Nucleic Acids Res 28:60-4.
- ↑ 5.0 5.1 5.2 CGSC: The Coli Genetics Stock Center
- ↑ 6.0 6.1 The Tn10 insertion sites determined by Nichols et al. 1998 (PMID:9829956) were reannotated by alignment with E. coli K-12 genome sequence (GenBank accession NC_000913). P1 contransduction frequencies were calculated using the formula of Wu (PMID:5338813).
- ↑ Zaslaver, A et al. (2006) A comprehensive library of fluorescent transcriptional reporters for Escherichia coli. Nat. Methods 3 623-8 PubMed
Categories
- Genes in OpenBioSystems with Promoter Fusions
- Genes with homologs in Apis mellifera
- Genes with homologs in Caenorhabditis briggsae
- Genes with homologs in Caenorhabditis elegans
- Genes with homologs in Canis familiaris
- Genes with homologs in Dictyostelium discoideum
- Genes with homologs in Drosophila melanogaster
- Genes with homologs in Drosophila pseudoobscura
- Genes with homologs in Homo sapiens
- Genes with homologs in Macaca mulatta
- Genes with homologs in Mus musculus
- Genes with homologs in Pan troglodytes
- Genes with homologs in Rattus norvegicus
- Genes with homologs in Takifugu rubripes
- Genes with homologs in Tetraodon nigroviridis
- Genes with homologs in Shigella flexneri
- Genes with homologs in E. coli O157


