ygeR:On One Page
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;} h2 .editsection { display:none;}</css>
| Quickview | Gene | Gene Product(s) | Expression | Evolution | On One Page |
| Quickview | | Gene | | Product(s) | | Expression| | Evolution | | References | |
Quickview
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
| Standard Name |
ygeR |
|---|---|
| Gene Synonym(s) |
ECK2861, b2865, JW2833[1], JW2833 |
| Product Desc. |
putative lipoprotein[2]; putative lipoprotein; predicted DNA-binding transcriptional regulator[3] Novel lipoprotein, function unknown, metalloprotease homolog; M23 family[4] |
| Product Synonyms(s) |
Tetratricopeptide repeat transcriptional regulator[1], B2865[2][1], YgeR[2][1] , ECK2861, JW2833, b2865 |
| Function from GO |
<GO_nr /> |
| Knock-Out Phenotype | |
| Regulation/Expression | |
| Regulation/Activity | |
| Quick Links | |
| edit table |
</protect> See Help:Quickview for help with entering information in the Quickview table. <protect></protect>
Notes
nlpD, ygeR, yebA and envC are paralogs. ygeR is a backbone gene flanking the insertion of the type three secretion system pathogenicity island ETT2. 18 kb of ETT2 has been deleted from K-12, leaving a 10 kb remnant. First 25 aa are a type II signal peptide.[4]
Gene
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
Nomenclature
See Help:Gene_nomenclature for help with entering information in the Gene Nomenclature table.
| Standard name |
ygeR |
|---|---|
| Mnemonic |
Systematic nomenclature |
| Synonyms |
ECK2861, b2865, JW2833[1], JW2833 |
| edit table |
</protect>
Notes
Location(s) and DNA Sequence
<protect>
See Help:Gene_location for help entering information in the Gene Location and DNA sequence table.
| Strain | Map location | Genome coordinates | Genome browsers | Sequence links |
|---|---|---|---|---|
|
MG1655 |
64.6 minutes |
MG1655: 2997913..2997158 |
||
|
NC_012967: 2885646..2884891 |
||||
|
NC_012759: 2884306..2885061 |
||||
|
W3110 |
|
W3110: 2998547..2997792 |
||
|
DH10B: 3091783..3091028 |
||||
| edit table |
</protect>
Notes
Sequence Features
See Help:Gene_sequence_features for help in entering sequence features in EcoliWiki.
| Feature Type | Strain | Genomic Location | Evidence | Reference | Notes |
|---|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Alleles and Phenotypes
See Help:Gene_alleles for how to enter or edit alleles and phenotypes in EcoliWiki.
| Allele | Nt change(s) | AA change(s) | Phenotype: Type | Phenotype: Description | Reference | Availability | Comments |
|---|---|---|---|---|---|---|---|
|
ΔygeR (Keio:JW2833) |
deletion |
deletion |
PMID:16738554 |
||||
|
ΔygeR787::kan |
PMID:16738554 |
||||||
|
ΔygeR |
deletion |
deletion |
Cell Shape |
defect in cell separation |
PMID:19525345 |
||
|
ΔygeR ΔenvC |
deletion |
deletion |
Cell Shape |
defect in cell separation |
PMID:19525345 |
||
|
ΔygeR ΔnlpD |
deletion |
deletion |
Cell Shape |
defect in cell separation |
PMID:19525345 |
||
|
ΔygeR ΔyebA |
deletion |
deletion |
Cell Shape |
defect in cell separation |
PMID:19525345 |
||
|
ΔygeR ΔnlpD ΔenvC |
deletion |
deletion |
Cell Shape |
defect in cell separation |
PMID:19525345 |
||
|
ΔygeR ΔyebA ΔenvC |
deletion |
deletion |
Cell Shape |
defect in cell separation |
PMID:19525345 |
||
|
ΔygeR ΔyebA ΔnlpD |
deletion |
deletion |
Cell Shape |
defect in cell separation |
PMID:19525345 |
||
|
ΔygeR ΔyebA ΔnlpD ΔenvC |
deletion |
deletion |
Cell Shape |
defect in cell separation |
PMID:19525345 |
| |
| edit table |
<protect></protect>
Notes
Genetic Interactions
<protect>
| Interactor | Interaction | Allele | Score(s) | Reference(s) | Accessions | Notes |
|---|---|---|---|---|---|---|
| edit table |
</protect>
Notes
Genetic Resources
See Help:Gene_resources for help entering information into the Genetic Resources table.
| Resource | Resource Type | Source | Notes/Reference |
|---|---|---|---|
|
JW2833 |
Plasmid clone |
PMID:16769691 Status:Clone OK Primer 1:GCCAGTGCGGGACGCCTGAATAA Primer 2:CCGCATTTTGGCTTGCTGCCCTG | |
|
Kohara Phage |
PMID:3038334 | ||
|
Kohara Phage |
PMID:3038334 | ||
|
Linked marker |
est. P1 cotransduction: 13% [6] | ||
|
Linked marker |
est. P1 cotransduction: % [6] | ||
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:G7484 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG13048 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120004047 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EchoBASE:EB2860 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0009407 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Product(s)
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Nomenclature
See Help:Product_nomenclature for help entering or editing information in this section of EcoliWiki.
| Standard name |
YgeR |
|---|---|
| Synonyms |
Tetratricopeptide repeat transcriptional regulator[1], B2865[2][1], YgeR[2][1] , ECK2861, JW2833, b2865 |
| Product description |
putative lipoprotein[2]; putative lipoprotein; predicted DNA-binding transcriptional regulator[3] Novel lipoprotein, function unknown, metalloprotease homolog; M23 family[4] |
| EC number (for enzymes) |
|
| edit table |
<protect></protect>
Notes
Function
<protect>
Gene Ontology
See Help:Gene_ontology for help entering or editing GO terms and GO annotations in EcoliWiki.
| Qualifier | GO ID | GO term name | Reference | Evidence Code | with/from | Aspect | Notes | Status |
|---|---|---|---|---|---|---|---|---|
|
GO:0000920 |
septum digestion after cytokinesis |
PMID:19525345 |
IGI: Inferred from Genetic Interaction |
UniProtKB:P0ADA3 UniProtKB:P37690 UniProtKB:P0AFS9
|
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0004222 |
metalloendopeptidase activity |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR002886 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005886 |
plasma membrane |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-1003 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006508 |
proteolysis |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR002886 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0000920 |
cytokinetic cell separation |
PMID:19525345 |
IGI: Inferred from Genetic Interaction |
UniProtKB:P37690
|
P |
complete | ||
|
GO:0016020 |
membrane |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0472 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0016998 |
cell wall macromolecule catabolic process |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR002482 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0000920 |
cytokinetic cell separation |
PMID:19525345 |
IGI: Inferred from Genetic Interaction |
UniProtKB:P0ADA3 UniProtKB:P37690 UniProtKB:Q46798
|
P |
complete | ||
|
GO:0016998 |
cell wall macromolecule catabolic process |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR018392 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0031225 |
anchored to membrane |
GO_REF:0000023 |
IEA: Inferred from Electronic Annotation |
SP_SL:SL-9901 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
Interactions See Help:Product_interactions for help entering or editing information about gene product interactions in this section of EcoliWiki.
| Partner Type | Partner | Notes | References | Evidence |
|---|---|---|---|---|
|
Protein |
rplL |
PMID:19402753 |
LCMS(ID Probability):99.6 | |
| edit table |
</protect>
Notes
Localization
See Help:Product_localization for how to add or edit information in this section of EcoliWiki.
| Compartment | Description | Evidence | Reference/Source | Notes | |
|---|---|---|---|---|---|
|
Outer membrane |
PMID:15174130 |
| |||
| edit table |
<protect></protect>
Notes
Structure and Physical Properties
<protect>
Physical Properties
See Help:Product_physical_properties for help entering or editing information about the physical properties of this gene product.
| Name | |
|---|---|
| Sequence |
MFFKRGKILS AGRLNKKSLG IVMLLSVGLL LAGCSGSKSS DTGTYSGSVY TVKRGDTLYR ISRTTGTSVK ELARLNGISP PYTIEVGQKL KLGGAKSSSI TRKSTAKSTT KTASVTPSSA VPKSSWPPVG QRCWLWPTTG KVIMPYSTAD GGNKGIDISA PRGTPIYAAG AGKVVYVGNQ LRGYGNLIMI KHSEDYITAY AHNDTMLVNN GQSVKAGQKI ATMGSTDAAS VRLHFQIRYR ATAIDPLRYL PPQGSKPKC |
| Length |
259 |
| Mol. Wt |
27.555 kDa |
| pI |
10.6 (calculated) |
| Extinction coefficient |
34,380 - 34,755 (calc based on 12 Y, 3 W, and 3 C residues) |
| edit table |
Domains/Motifs/Modification Sites
|
See Help:Product_domains_motifs for help entering or editing information in this section of EcoliWiki.
|
<motif_map/> |
|
Structure
| </protect>
Structure figures<protect> | ||||||
Notes
Gene Product Resources
See Help:Product_resources for help with entering or editing information in this section of EcoliWiki.
| Resource type | Source | Notes/Reference |
|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
ASAP:ABE-0009407 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EcoCyc:G7484 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG13048 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
RegulonDB:ECK120004047 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB2860 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Expression
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Overview
This is a placeholder for a summary statement on how expression of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Cellular Levels
| Molecule | Organism or Strain | Value | Units | Experimental Conditions | Assay used | Notes | Reference(s) |
|---|---|---|---|---|---|---|---|
|
Protein |
E. coli K-12 MG1655 |
298 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
125 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
262 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
| edit table |
Notes
Transcription and Transcriptional Regulation
<protect>
|
See Help:Expression_transcription for help entering or editing information in this section of EcoliWiki.
|
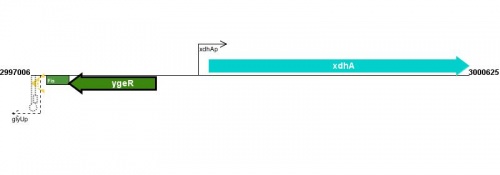 Figure courtesy of RegulonDB |
</protect>
Notes
This is a placeholder for a summary statement about how transcription of this gene is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Translation and Regulation of Translation
<protect><gbrowseImage>
name=NC_000913:2997893..2997933
source=MG1655
flip=1
type=Gene+DNA_+Protein
preset=Nterminus
</gbrowseImage>
This picture shows the sequence around the N-terminus.
</protect>
Notes
This is a placeholder for a summary statement about how translation of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Turnover and Regulation of Turnover
</protect>
Notes
This is a placeholder for a summary statement about turnover of this gene product. You can help EcoliWiki by becoming a user and writing/editing this statement.
Experimental
<protect>
Mutations Affecting Expression
See Help:Expression_mutations for help entering or editing information in this section of EcoliWiki.
| Allele Name | Mutation | Phenotype | Reference |
|---|---|---|---|
| edit table |
Expression Studies
See Help:Expression_studies for help entering or editing information in this section of EcoliWiki.
| Type | Reference | Notes |
|---|---|---|
|
microarray |
NCBI GEO profiles for ygeR | |
|
microarray |
Summary of data for ygeR from multiple microarray studies | |
| edit table |
Expression Resources
See Help:Expression_resources for help entering or editing information in this section of EcoliWiki.
| Resource Name | Resource Type | Description | Source | Notes |
|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions Related to ygeR Expression
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:G7484 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB2860 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG13048 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120004047 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0009407 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Evolution
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Homologs in Other Organisms
See Help:Evolution_homologs for help entering or editing information in this section of EcoliWiki.
| Organism | Homologs (Statistics) | Comments |
|---|---|---|
|
Shigella flexneri |
S3067 |
From SHIGELLACYC |
|
E. coli O157 |
Z4203 |
From ECOO157CYC |
| edit table |
Do-It-Yourself Web Tools
<protect></protect>
Notes
Families
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:G7484 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG13048 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120004047 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB2860 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0009407 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
References
See Help:References for how to manage references in EcoliWiki.
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 Riley, M. et al. (2006) Nucleic Acids Res 34:1-6 (corrected supplemental data from B. Wanner)
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 2.6 EcoCyc (release 10.6; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 3.0 3.1 EcoCyc (release 11.1; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 4.0 4.1 4.2 EcoGene: Rudd, KE (2000) EcoGene: a genome sequence database for Escherichia coli K-12. Nucleic Acids Res 28:60-4.
- ↑ 5.0 5.1 5.2 CGSC: The Coli Genetics Stock Center
- ↑ 6.0 6.1 The Tn10 insertion sites determined by Nichols et al. 1998 (PMID:9829956) were reannotated by alignment with E. coli K-12 genome sequence (GenBank accession NC_000913). P1 contransduction frequencies were calculated using the formula of Wu (PMID:5338813).
Categories


