tilS:On One Page
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;} h2 .editsection { display:none;}</css>
| Quickview | Gene | Gene Product(s) | Expression | Evolution | On One Page |
| Quickview | | Gene | | Product(s) | | Expression| | Evolution | | References | |
Quickview
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
| Standard Name |
tilS |
|---|---|
| Gene Synonym(s) |
ECK0187, b0188, JW0183, mesJ, yaeN[1], yaeN |
| Product Desc. |
tRNA(Ile) lysidine (L34) synthase, ATP-dependent; solely responsible for tRNA(Ile) lysidine 34 (L34) formation[4] |
| Product Synonyms(s) |
tRNA(Ile)-lysidine synthetase[1], B0188[2][1], YaeN[2][1], MesJ[2][1], TilS[2][1] , ECK0187, JW0183, mesJ, yaeN, b0188 |
| Function from GO |
<GO_nr /> |
| Knock-Out Phenotype | |
| Regulation/Expression | |
| Regulation/Activity | |
| Quick Links | |
| edit table |
</protect> See Help:Quickview for help with entering information in the Quickview table. <protect></protect>
Notes
YaeN (TilS) is mistakenly labeled as a cell cycle gene in many databases. the confusion arose from the misinterpretation of Genbank Z50870. the real "minE suppressor" is the small rof gene next to tilS, labeled as orf4 in Z50870. PDB: 1NI5. Gu M., Burling T., Lima C.D., Structure of the MesJ PP-ATPase from Escherichia coli, unpublished. tilS is an essential gene. AFMB Structural Genomics target No. 121 (http://afmb.cnrs-mrs.fr/article171.html).[4]
Gene
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
<protect>
Nomenclature
See Help:Gene_nomenclature for help with entering information in the Gene Nomenclature table.
| Standard name |
tilS |
|---|---|
| Mnemonic |
tRNA(Ile) lysidine synthetase |
| Synonyms |
ECK0187, b0188, JW0183, mesJ, yaeN[1], yaeN |
| edit table |
</protect>
Notes
Location(s) and DNA Sequence
<protect>
See Help:Gene_location for help entering information in the Gene Location and DNA sequence table.
| Strain | Map location | Genome coordinates | Genome browsers | Sequence links |
|---|---|---|---|---|
|
MG1655 |
4.58 minutes |
MG1655: 212331..213629 |
||
|
NC_012967: 215173..216471 |
||||
|
NC_012759: 212330..213628 |
||||
|
W3110 |
|
W3110: 212331..213629 |
||
|
DH10B: 186435..187733 |
||||
| edit table |
</protect>
Notes
Sequence Features
See Help:Gene_sequence_features for help in entering sequence features in EcoliWiki.
| Feature Type | Strain | Genomic Location | Evidence | Reference | Notes |
|---|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Alleles and Phenotypes
See Help:Gene_alleles for how to enter or edit alleles and phenotypes in EcoliWiki.
| Allele | Nt change(s) | AA change(s) | Phenotype: Type | Phenotype: Description | Reference | Availability | Comments |
|---|---|---|---|---|---|---|---|
|
mesJ |
Nucleoside synthesis |
lysidine synthesis is defective in about 50% of the tRNAlle2 molecules. See figure 3B. |
PMID:14527414 |
lysidine synthesis is defective in about 50% of the tRNAlle2 molecules. See figure 3B. | |||
| edit table |
<protect></protect>
Notes
Genetic Interactions
<protect>
| Interactor | Interaction | Allele | Score(s) | Reference(s) | Accessions | Notes |
|---|---|---|---|---|---|---|
| edit table |
</protect>
Notes
Genetic Resources
See Help:Gene_resources for help entering information into the Genetic Resources table.
| Resource | Resource Type | Source | Notes/Reference |
|---|---|---|---|
|
JW0183 |
Plasmid clone |
PMID:16769691 Status:Clone OK Primer 1:GCCACACTCACGCTCAATAGACA Primer 2:CCACTAAGCGTTTTCTGCCAGAC | |
|
Kohara Phage |
PMID:3038334 | ||
|
Kohara Phage |
PMID:3038334 | ||
|
Linked marker |
est. P1 cotransduction: 3% [6] | ||
|
Linked marker |
est. P1 cotransduction: 49% [6] | ||
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:G6096 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG13220 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
RegulonDB:ECK120002716 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB3011 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0000638 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Product(s)
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Nomenclature
See Help:Product_nomenclature for help entering or editing information in this section of EcoliWiki.
| Standard name |
TilS |
|
|---|---|---|
| Synonyms |
tRNA(Ile)-lysidine synthetase[1], B0188[2][1], YaeN[2][1], MesJ[2][1], TilS[2][1] , ECK0187, JW0183, mesJ, yaeN, b0188 |
|
| Product description |
tRNA(Ile) lysidine (L34) synthase, ATP-dependent; solely responsible for tRNA(Ile) lysidine 34 (L34) formation[4] |
|
| EC number (for enzymes) |
| |
| edit table |
<protect></protect>
Notes
Function
<protect>
Gene Ontology
See Help:Gene_ontology for help entering or editing GO terms and GO annotations in EcoliWiki.
| Qualifier | GO ID | GO term name | Reference | Evidence Code | with/from | Aspect | Notes | Status |
|---|---|---|---|---|---|---|---|---|
|
GO:0000166 |
nucleotide binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012094 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0000166 |
nucleotide binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012795 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006400 |
tRNA modification |
PMID:14527414 |
IDA: Inferred from Direct Assay |
P |
Purified recombinant TilS protein creates the modified nucleoside lysidine in an in vitro reaction with ATP, lysine, and purified precursor tRNAIle2. |
complete | ||
|
GO:0000166 |
nucleotide binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012796 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0032267 |
tRNA(Ile)-lysidine synthase activity |
PMID:14527414 |
IDA: Inferred from Direct Assay |
F |
Purified recombinant TilS protein creates the modified nucleoside lysidine in an in vitro reaction with ATP, lysine, and purified precursor tRNAIle2. |
complete | ||
|
GO:0000166 |
nucleotide binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR015262 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0000166 |
nucleotide binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0547 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012094 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012795 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012796 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR015262 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005524 |
ATP binding |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0067 |
F |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005737 |
cytoplasm |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012094 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005737 |
cytoplasm |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012795 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005737 |
cytoplasm |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012796 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005737 |
cytoplasm |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR015262 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005737 |
cytoplasm |
GOA:spkw |
IEA: Inferred from Electronic Annotation |
SP_KW:KW-0963 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0005737 |
cytoplasm |
GO_REF:0000023 |
IEA: Inferred from Electronic Annotation |
SP_SL:SL-0086 |
C |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006400 |
tRNA modification |
PMID:14527414 |
IDA: Inferred from Direct Assay |
P |
Seeded from EcoCyc (v14.0) |
complete | ||
|
GO:0006412 |
translation |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012094 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006412 |
translation |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012795 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006412 |
translation |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR012796 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
|
GO:0006412 |
translation |
GOA:interpro |
IEA: Inferred from Electronic Annotation |
InterPro:IPR015262 |
P |
Seeded from EcoCyc (v14.0) |
complete | |
| edit table |
Interactions See Help:Product_interactions for help entering or editing information about gene product interactions in this section of EcoliWiki.
| Partner Type | Partner | Notes | References | Evidence |
|---|---|---|---|---|
|
RNA |
tRNAIle2 |
Purified TilS binding to tRNAIle2 detected by gel shift analysis [7]. |
||
|
Protein |
rplA |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
rpsI |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
moaC |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
rplO |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
rpsG |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
fur |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
rplJ |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
frdC |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
dinG |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
uidC |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
rpsA |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
rplQ |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
bglX |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
|
Protein |
rplB |
PMID:16606699 |
Experiment(s):EBI-1135871 | |
| edit table |
</protect>
Notes
Localization
See Help:Product_localization for how to add or edit information in this section of EcoliWiki.
| Compartment | Description | Evidence | Reference/Source | Notes |
|---|---|---|---|---|
| edit table |
<protect></protect>
Notes
Structure and Physical Properties
<protect>
Physical Properties
See Help:Product_physical_properties for help entering or editing information about the physical properties of this gene product.
| Name | |
|---|---|
| Sequence |
MTLTLNRQLL TSRQILVAFS GGLDSTVLLH QLVQWRTENP GVALRAIHVH HGLSANADAW VTHCENVCQQ WQVPLVVERV QLAQEGLGIE AQARQARYQA FARTLLPGEV LVTAQHLDDQ CETFLLALKR GSGPAGLSAM AEVSEFAGTR LIRPLLARTR GELVQWARQY DLRWIEDESN QDDSYDRNFL RLRVVPLLQQ RWPHFAEATA RSAALCAEQE SLLDELLADD LAHCQSPQGT LQIVPMLAMS DARRAAIIRR WLAGQNAPMP SRDALVRIWQ EVALAREDAS PCLRLGAFEI RRYQSQLWWI KSVTGQSENI VPWQTWLQPL ELPAGLGSVQ LNAGGDIRPP RADEAVSVRF KAPGLLHIVG RNGGRKLKKI WQELGVPPWL RDTTPLLFYG ETLIAAAGVF VTQEGVAEGE NGVSFVWQKT LS |
| Length |
432 |
| Mol. Wt |
48.203 kDa |
| pI |
3.5 (calculated) |
| Extinction coefficient |
89,950 - 90,700 (calc based on 5 Y, 15 W, and 6 C residues) |
| edit table |
Domains/Motifs/Modification Sites
|
See Help:Product_domains_motifs for help entering or editing information in this section of EcoliWiki.
|
<motif_map/> |
|
Structure
| </protect>
Structure figures<protect> | ||||||
Notes
Gene Product Resources
See Help:Product_resources for help with entering or editing information in this section of EcoliWiki.
| Resource type | Source | Notes/Reference |
|---|---|---|
| edit table |
<protect></protect>
Notes
Accessions in Other Databases
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
ASAP:ABE-0000638 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
EcoCyc:G6096 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG13220 |
Escherichia coli str. K-12 substr. MG1655 | |
|
Escherichia coli str. K-12 substr. MG1655 | ||
|
RegulonDB:ECK120002716 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB3011 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Expression
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Overview
This is a placeholder for a summary statement on how expression of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Cellular Levels
| Molecule | Organism or Strain | Value | Units | Experimental Conditions | Assay used | Notes | Reference(s) |
|---|---|---|---|---|---|---|---|
|
Protein |
E. coli K-12 MG1655 |
490 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
122 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
|
Protein |
E. coli K-12 MG1655 |
221 |
molecules/cell/generation |
|
Ribosome Profiling |
PMID: 24766808 | |
| edit table |
Notes
Transcription and Transcriptional Regulation
<protect>
|
See Help:Expression_transcription for help entering or editing information in this section of EcoliWiki.
|
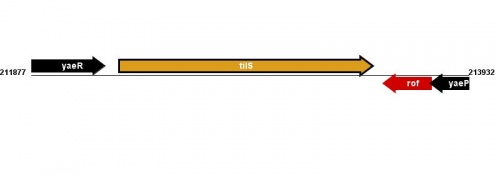 Figure courtesy of RegulonDB |
</protect>
Notes
This is a placeholder for a summary statement about how transcription of this gene is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Translation and Regulation of Translation
<protect><gbrowseImage>
name=NC_000913:212311..212351
source=MG1655
type=Gene+DNA_+Protein
preset=Nterminus
</gbrowseImage>
This picture shows the sequence around the N-terminus.
</protect>
Notes
This is a placeholder for a summary statement about how translation of this gene product is regulated. You can help EcoliWiki by becoming a user and writing/editing this statement.
Turnover and Regulation of Turnover
</protect>
Notes
This is a placeholder for a summary statement about turnover of this gene product. You can help EcoliWiki by becoming a user and writing/editing this statement.
Experimental
<protect>
Mutations Affecting Expression
See Help:Expression_mutations for help entering or editing information in this section of EcoliWiki.
| Allele Name | Mutation | Phenotype | Reference |
|---|---|---|---|
| edit table |
Expression Studies
See Help:Expression_studies for help entering or editing information in this section of EcoliWiki.
| Type | Reference | Notes |
|---|---|---|
|
microarray |
NCBI GEO profiles for tilS | |
|
microarray |
Summary of data for tilS from multiple microarray studies | |
| edit table |
Expression Resources
See Help:Expression_resources for help entering or editing information in this section of EcoliWiki.
| Resource Name | Resource Type | Description | Source | Notes |
|---|---|---|---|---|
|
GFP Fusion |
Intergenic region (212181..212401) fused to gfpmut2. |
GFP fusion described in Zaslaver, et al. | ||
| edit table |
<protect></protect>
Notes
Accessions Related to tilS Expression
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:G6096 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB3011 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG13220 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120002716 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0000638 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
Evolution
{{#css:Suppresslinks.css}}<css>h1 .editsection { display:none;}
h2 .editsection { display:none;}</css>
Homologs in Other Organisms
See Help:Evolution_homologs for help entering or editing information in this section of EcoliWiki.
| Organism | Homologs (Statistics) | Comments |
|---|---|---|
|
Schizosaccharomyces pombe |
|
From Inparanoid:20070104 |
|
Shigella flexneri |
MESJ |
From SHIGELLACYC |
|
E. coli O157 |
MESJ |
From ECOO157CYC |
| edit table |
Do-It-Yourself Web Tools
<protect></protect>
Notes
Families
See Help:Gene_accessions for help with entering information into the Gene Accessions table.
| Database | Accession | Notes |
|---|---|---|
|
EcoCyc:G6096 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EcoGene:EG13220 |
Escherichia coli str. K-12 substr. MG1655 | |
|
RegulonDB:ECK120002716 |
Escherichia coli str. K-12 substr. MG1655 | |
|
EchoBASE:EB3011 |
Escherichia coli str. K-12 substr. MG1655 | |
|
ASAP:ABE-0000638 |
Escherichia coli str. K-12 substr. MG1655 | |
| edit table |
<protect></protect>
Notes
Links
| Name | URL | Comments |
|---|---|---|
| edit table |
References
See Help:References for how to manage references in EcoliWiki.
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 Riley, M. et al. (2006) Nucleic Acids Res 34:1-6 (corrected supplemental data from B. Wanner)
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 EcoCyc (release 10.6; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 3.0 3.1 EcoCyc (release 11.1; 2007) Keseler, IM et al. (2005) Nucleic Acids Res. 33(Database issue):D334-7
- ↑ 4.0 4.1 4.2 EcoGene: Rudd, KE (2000) EcoGene: a genome sequence database for Escherichia coli K-12. Nucleic Acids Res 28:60-4.
- ↑ 5.0 5.1 CGSC: The Coli Genetics Stock Center
- ↑ 6.0 6.1 The Tn10 insertion sites determined by Nichols et al. 1998 (PMID:9829956) were reannotated by alignment with E. coli K-12 genome sequence (GenBank accession NC_000913). P1 contransduction frequencies were calculated using the formula of Wu (PMID:5338813).
- ↑ Soma, A et al. (2003) An RNA-modifying enzyme that governs both the codon and amino acid specificities of isoleucine tRNA. Mol. Cell 12 689-98 PubMed
- ↑ Zaslaver, A et al. (2006) A comprehensive library of fluorescent transcriptional reporters for Escherichia coli. Nat. Methods 3 623-8 PubMed
Categories


