Many Microbe Microarrays Database (M3D)
You can help EcoliWiki by editing the content of this page. For information about becoming a registered user and obtaining editing privileges, see Help:Accounts. <protect>
| Link/URL: |
Many Microbe Microarrays Database Mirror site: http://cgs.wustl.edu/~faithj/m3d_mirror/ |
|---|---|
| What: |
Affymetrix data for E. coli and other bacteria |
| Who: |
Tim Gardner http://gardnerlab.bu.edu/ Boston University |
| Updates: | |
| Upcoming events: | |
| Web Services: |
|
| edit table |
</protect>
Contents
About Many Microbe Microarrays Database (M3D)
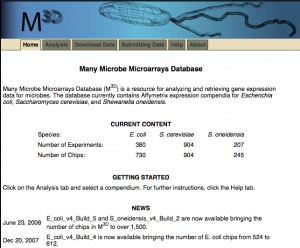
Many Microbe Microarrays Database (M3D) is a resource for analyzing and retrieving gene expression data for microbes. The database currently contains affymetrix expression profiles for Escherichia coli. In the future, Shewanella oneidensis affymetrix data will be available as well.[1]
Content
Experiments: 189 Arrays: 445
Using Many Microbe Microarrays Database (M3D)
Browsing
Searching
Users can query M3D by choosing a project and a list of genes. Data can be normalized (scaling by gene) prior to analysis.
http://m3d.bu.edu/cgi-bin/web/array/index.pl?db=E_coli_v3_Build_1
Presentation formats include:
- Heat Plot (genes vs experiments)
- Histogram of individual gene expression (one plot per gene)
- Histogram of all gene expression (one plot for N genes)
- Plot Expression of many genes as a function of one gene
- Link to download expression data
Average Array Replicates by:
- RMA
- GCRMA
- DchipPM
- DchipMM
- MAS5
Heat Plot Parameters:
- rescale data to within stddev of mean
- Order Genes By Correlation
- Order Experiments By Correlation
Heat Map coloring:
- Red/Blue
- Red/Green
M3D provides a meatalink to execute queries on N-browse, which is described as an interactive graphical browser for molecular interaction networks
http://nematoda.bio.nyu.edu:8080/NBrowse/N-Browse.jsp?last=false
Usage examples
Add links to additional pages describing success stories here.
Technology
The following free software and libraries were used for array normalization and to develop M3D: dChip, Bioconductor (for GCRMA and RMA), gd, cgic, Cluster 3.0, and MySQL.
Web Services/API
Discussion
References
See Help:References for how to manage references in EcoliWiki.
External Links
Many Microbe Microarrays Database (M3D)